MERCURIUS™
Blood BRB-seq
Library preparation kits for Illumina®
96, 384 RNA-sea library preps in one tube.
| Catalog number | #10823 | #11023 | #10825 | #11025 |
| Total preps
|
96 | 384 | 384 | 1'536 |
| RNA and plate multiplexing format
|
96 | 96 | 384 | 384 |
| Barcoded oligo-dT plates included
| 1 | 4 | 1 | 4 |
| UDI pairs included
|
4 | 4 | 4 | 4 |
| Globin blockers | included | included | included | included |
| Documentation | ||||
| #10,823 Cat ##10823 | ||||
|---|---|---|---|---|
| Total reactions
|
||||
| RNA multiplexing format
|
96 | |||
| UDI pairs included
|
4 | |||
| Globin blockers | included | |||
| Cat ##11023 | ||||
|---|---|---|---|---|
| Total reactions
|
384 | |||
| RNA multiplexing format
|
96 | |||
| UDI pairs included
|
4 | |||
| Globin blockers | included | |||
| Cat ##10825 | ||||
|---|---|---|---|---|
| Total reactions
|
384 | |||
| RNA multiplexing format
|
384 | |||
| UDI pairs included
|
4 | |||
| Globin blockers | included | |||
| Cat ##11025 | ||||
|---|---|---|---|---|
| Total reactions
|
||||
| RNA multiplexing format
|
384 | |||
| UDI pairs included
|
4 | |||
| Globin blockers | included | |||
Add-ons
Accessories
UDI expansion
module

- Compatible with the MERCURIUS™ product line.
- 12 UDI pairs.
- Flexible dual indexing solutions.
Package
Libraries
sequencing
$3
per million reads
- Take advantage of our low sequencing price, which is enabled by the recurrent sequencing runs.
- Your libraries will be sequenced in large sequencing runs with other compatible BRB-seq and DRUG-seq libraries.
Package
Data
pre-processing
$600 NOW $150
per 96 samples
- We provide you with everything from raw data to gene count matrices and sequencing report.
- Guaranteed turn-on time: 2 days.
-
Benefits
The MERCURIUS™ Blood BRB-seq library preparation kits for Illumina® contain all the oligos and enzymes needed to go from purified blood RNA to sequencing-ready DNA libraries.
Bulk RNA sequencingat scaleIntegratedglobin depletionOne-day labworkflowMore samples, more replicates. Robust results, significant discoveries.
Seamlessly integrated globin depletion. No need to purchase additional kits.
Convenient and short protocol from samples to sequencing-ready libraries in one day.
-
Experimental workflow at a glance
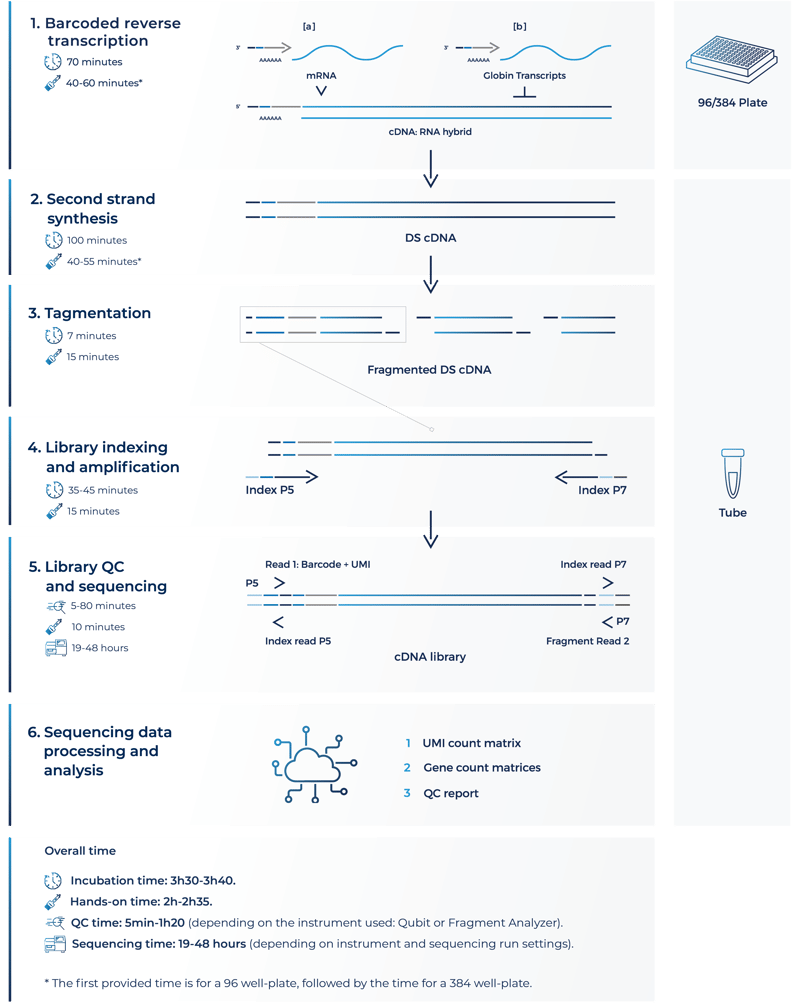
-
Performance
1. Uniform detection of 15'000+ genes at 1 million reads per sample across 96 samples
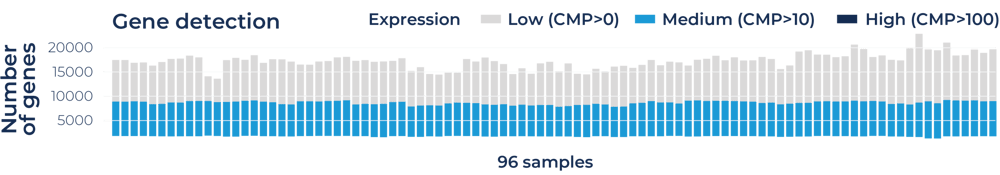
Distribution of the number of detected genes across 96 blood RNA samples prepared with the MERCURIUS™ Blood BRB-seq library preparation kit. The library was sequenced at an average of 1 million reads per sample on an Illumina NextSeq 550.
2. The Blood BRB-seq provides state-of-the-art performance in terms of globin depletion efficiency
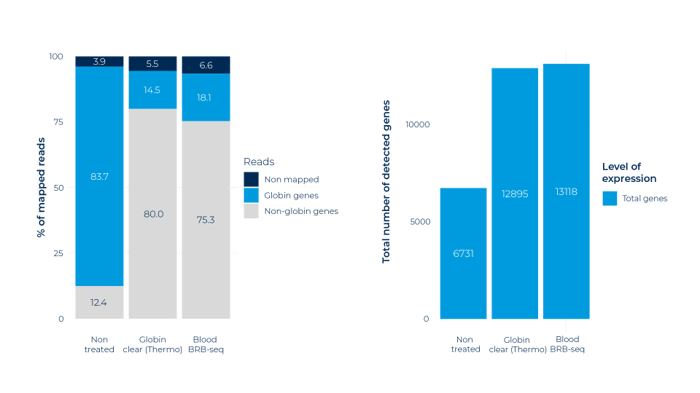
(left) Proportion of mapped reads assigned to globin genes (blue) and other genes (grey) for blood RNA samples without globin depletion and with globin depletion performed using ThermoFisher’s GLOBINclear™-Human Kit and the Blood BRB-seq method. (right) Number of detected genes across the three different conditions. Non-depleted libraries show a sharp decrease in performance, while BRB-seq and Globin Clear libraries both provide rich globin-free data.
3. Globin depletion is effective and specific to the target globin genes
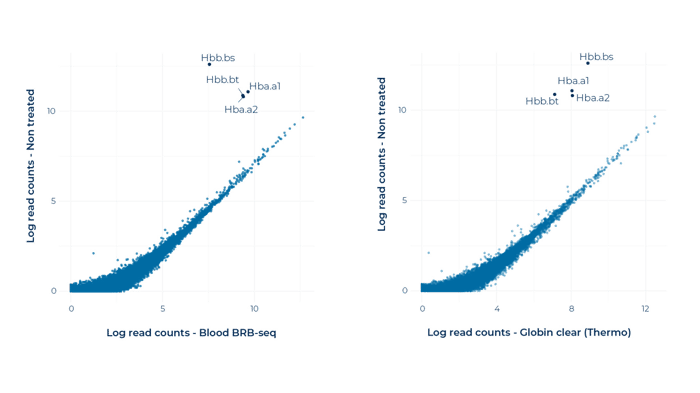
Correlation plots between mapped reads in between non-depleted libraries and Blood BRB-seq (left) and Globin Clear-depleted libraries (right). For both methods, globin depletion is effective and specific to the target globin genes, with negligible off-target effects on other genes.
4. Blood BRB-seq provides effective and high-throughput inline depletion of globin genes
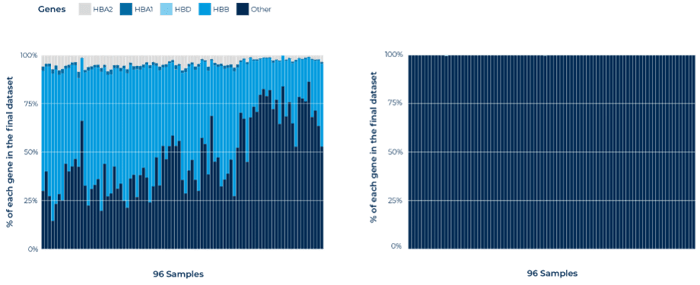
(left) Read distribution to the main globin genes on BRB-seq libraries WITHOUT globin depletion. The majority of mapped genes are associated with globin genes. (right) Read distribution to the main globin genes on BRB-seq libraries WITH globin depletion. The reads associated to globin genes are drastically reduced.
-
Product specifications
For (application)
3' mRNA sequencing
For use with (equipment)
Illumina NGS instruments
Species compatibility
Human and mouse
Available formats
96, 384, 1'536 reactions
Shipping conditions
Dry ice
Storage conditions
-20C
-

Human
Number of samples:96Reads per sample in demo dataset:10'000 readsTo have access to the deep-sequenced dataset (4.8M reads per sample) contact us.Demo dataset file size:57.7 MB
FAQs
-
Each BRB-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different BRB-seq pools.
To note, the total number of RNA samples that can be processed with one kit does not exceed the kit specifications; for instance, a 96-samples kit can be used to prepare up-to 96 samples distributed across up-to four different libraries.
-
The recommended range of RNA amount for each sample is of 50ng-1μg, normally the more RNA, the better.
The minimum recommended RIN number is 6 and the A260/230 ratio (Nanodrop) should be in the 1.5-2.2 range.
-
The only difference between BRB-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the BRB-seq barcode sequence.
For a thorough description of BRB-seq data processing, please refer to the BRB-seq kit user guide.
-
One of the key advantages of BRB-seq is that it does not only save reagents and cost in the library preparation stage, but also in the sequencing one.
As opposed to standard RNA-seq, where 20M-30M reads per sample are required, we normally recommend to sequence BRB-seq libraries at a depth of 4M-5M reads per sample, which is normally enough to detect the vast majority of expressed genes.
-
The barcode set for your kit is conveniently located on the kit label. Please refer to the label for accurate identification.
For optimal compatibility, ensure that you use the appropriate plate format (e.g., for kits designed for 96 reactions, the 96 well-plate format should be used). This ensures accurate and efficient processing of your samples. If you have any further questions or concerns, please contact our support team for assistance by email or using our live chat tool.
Speak with our RNA sequencing experts
Book a one-on-one call with one of our RNA experts to discover how we can assist your next project.