Discover Sample Datasets
Human
Built for teams aiming to detect full-length protein coding transcripts, isoforms, and splice variants in purified RNA from larger cohorts, more conditions, or more timepoints than possible with standard total RNA-seq methods.
MERCURIUS™ Full-Length BRB-seq combines early sample barcoding and multiplexing for full-length coverage of coding transcripts for up to 96 purified RNA samples in a single tube. It enables high-throughput mRNA-seq on Illumina® and AVITI™ platforms without compromising depth, data quality, or sensitivity compared to sample-by-sample methods.

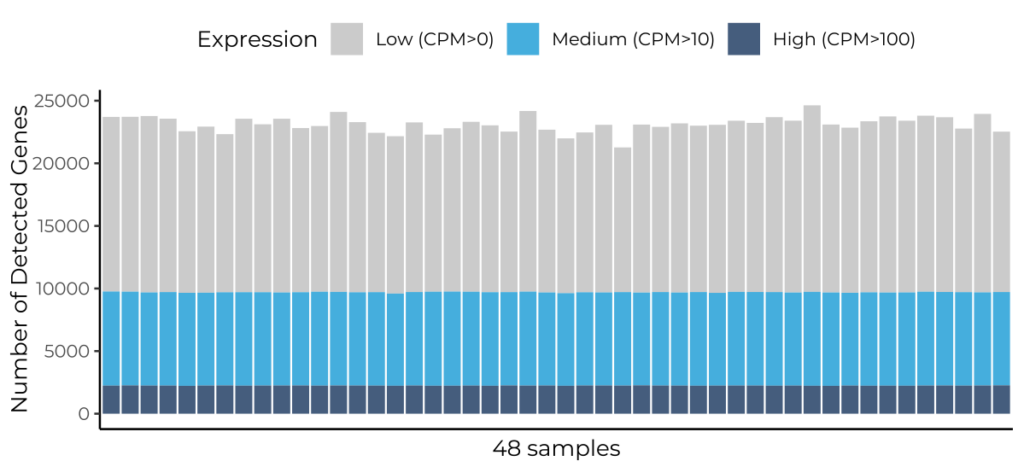
Number of genes detected per sample across a 48-sample MERCURIUS™ Full-Length BRB-seq run, stratified by expression level. All 48 samples show highly uniform gene detection, with approximately 22,000–24,000 genes detected at 12M reads/sample. The consistency across samples demonstrates the reproducibility of MERCURIUS™ Full-Length BRB-seq for high-throughput transcriptomic profiling.
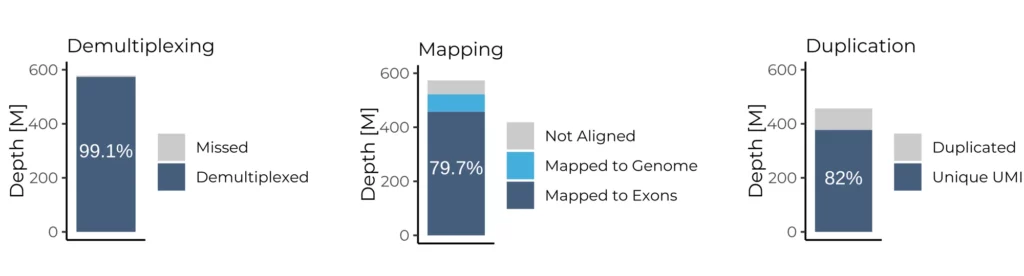
Sequencing quality metrics across three data preprocessing stages for a MERCURIUS™ Full-Length BRB-seq library. (Left) 99.1% of reads were successfully demultiplexed, confirming highly accurate barcode assignment with minimal read loss. (Centre) 79.7% of mapped reads aligned to exonic regions, indicating efficient transcript capture. (Right) 82% of reads represent unique UMI-tagged molecules, indicating low duplication rates and high library complexity. Together, these metrics demonstrate the technical robustness and sequencing efficiency of MERCURIUS™ Full-Length BRB-seq.
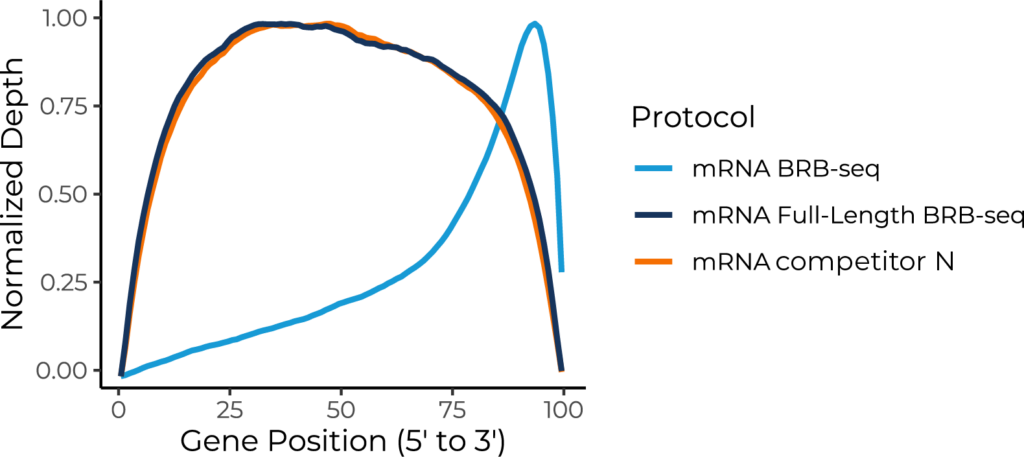
Normalized read depth across gene bodies (5′ to 3′). MERCURIUS™ Full-Length BRB-seq produces a uniform coverage profile across the entire gene body length, closely matching Competitor N. In contrast, standard BRB-seq shows characteristic 3′-end enrichment, owing to its 3′-capture design. This illustrates the advantage of MERCURIUS™ Full-Length BRB-seq for applications requiring uniform transcript coverage, such as isoform detection, splice variant analysis, and allele-specific expression.
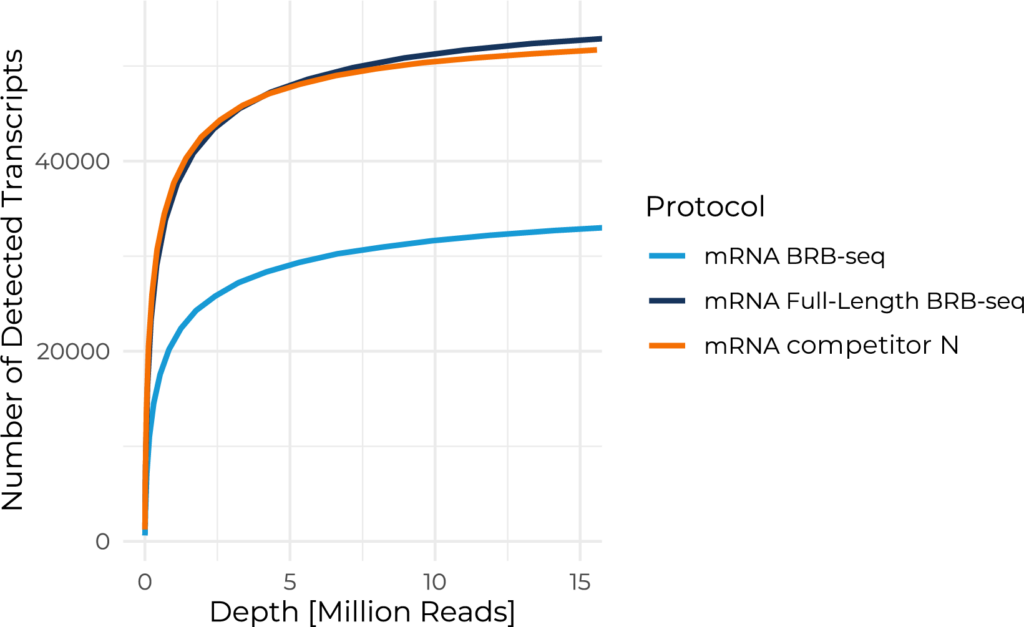
Number of detected transcripts as a function of downsampled sequencing depth for different technologies. MERCURIUS™ Full-Length BRB-seq closely tracks Competitor N across all read depths. Standard BRB-seq detects fewer transcripts, reflecting its 3′-capture design, which limits isoform resolution. These results demonstrate that MERCURIUS™ Full Length BRB-seq delivers competitor-equivalent transcript detection at higher throughput and lower cost.
Full-Length (Plant) BRB-seq performance shows 99% demultiplexing rate from raw data, 78% mapping rate, and 18% duplication rate of the 48 pooled samples and sequenced at 12 million reads per sample.
The gene body coverage shows a consistent and uniform reads distribution across the entire gene body for the Full-Length (Plant) BRB-seq protocol, comparable to the NEB protocol, while the BRB-seq protocol shows a significant 3′ bias due to its poly-A selection methodology.
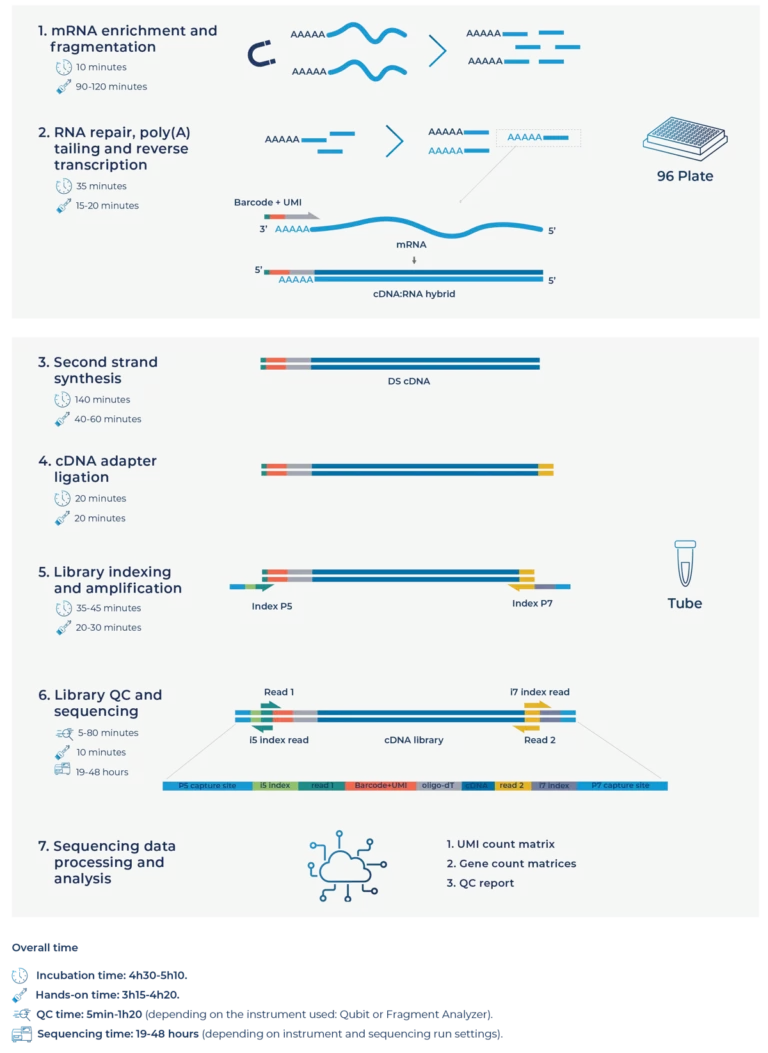
To have access to the deep-sequenced dataset, contact us.
Each Full-Length BRB-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different BRB-seq pools.
For instance, the 96-samples kit can be used to prepare up-to 96 samples distributed across up-to four different libraries.
The recommended range of RNA amount for each sample is of 10-1000 ng, normally the more RNA, the better.
The minimum recommended RIN number is 6 and the A260/230 ratio (Nanodrop) should be in the 1.5-2.2 range.
The only difference between Full-Length BRB-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the BRB-seq barcode sequence.
For a thorough description of Full-Length BRB-seq data processing, please look at the BRB-seq kit user guide.
The barcode set for your kit is conveniently located on the kit label. Please refer to the label for accurate identification.
For optimal compatibility, ensure that you use the appropriate plate format (e.g., for kits designed for 96 reactions, the 96 well-plate format should be used). This ensures accurate and efficient processing of your samples. If you have any further questions or concerns, please contact our support team for assistance by email or using our live chat tool.
Full-Length BRB-seq provides comprehensive coverage of the full-length mRNA transcripts.
We, therefore, normally recommend sequencing 12-20 million reads for each sample, which enables the reliable and unbiased detection of over 20,000 genes.
Product
MERCURIUS™ UDI X-Leap Expansion module
MERCURIUS™ Full-Length Post-Pooling Preparation Module (4 libraries)
Each kit includes 4 UDI pairs. Add the expansion module if you need more unique indexes (total 16 UDI pairs available).
Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…