
Capture expression, splicing, and isoforms, coding and non-coding RNAs.
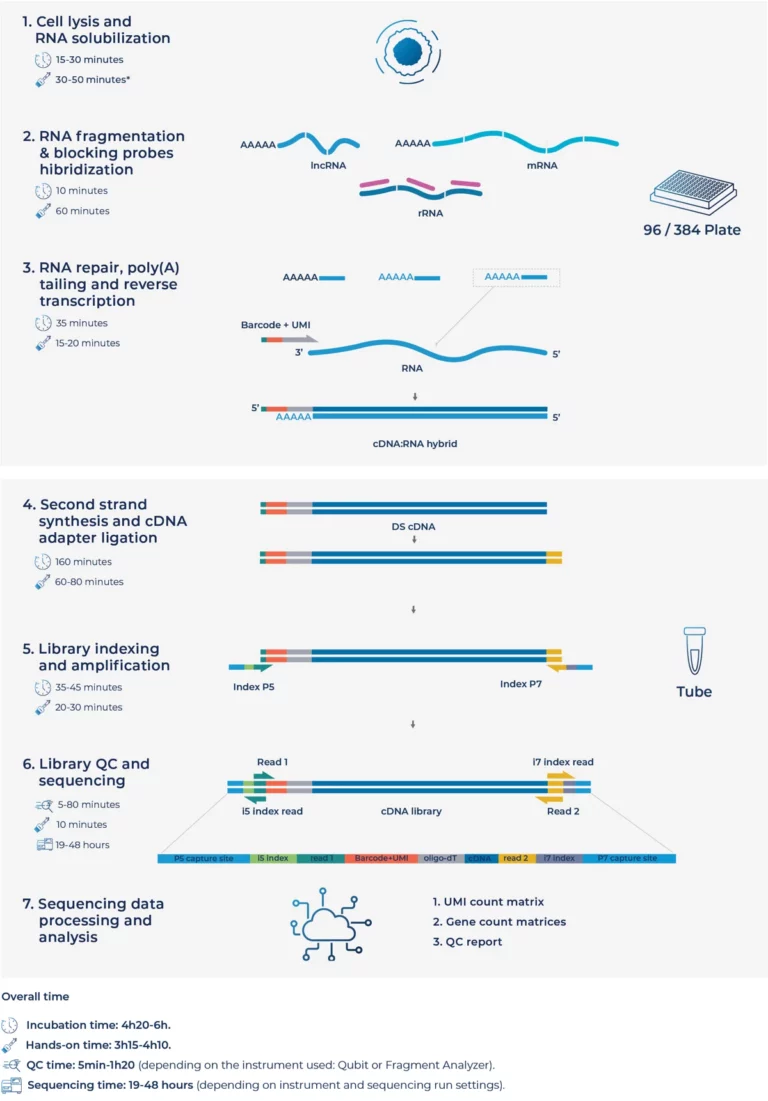
Process cells directly using optimized in-well lysis and reverse transcription.
Achieve higher mapping and gene detection rates with an improved workflow.
Generate sequencing-ready libraries from cells in a single working day.
Built for screening teams requiring high-throughput detection of coding and non-coding transcripts, isoforms, and splice variants in 2D cell lines and primary cells without RNA extraction. Ideal for studies with larger cohorts, more conditions, or more timepoints than possible to assess with standard total RNA-seq methods.
MERCURIUS™ Total DRUG-seq enables total RNA-seq at screening scale through in-well cell lysis, early barcoding, and multiplexing of 96 or 384 samples in a single tube. It provides full-length transcript coverage of coding and non-coding transcripts without compromising depth, data quality, or sensitivity compared to sample-by-sample methods. It enables ultra-high-throughput total RNA-seq on Illumina® and AVITI™ platforms and is compatible with standard high-throughput screening infrastructure.
Total RNA bulk RNA-seq for compound and dose–response screens where non-coding RNAs and full-length transcripts are informative.
Mechanism-of-action and toxicity profiling with coding and non-coding RNA signatures from cell-based assays.
Generating AI/ML-ready whole-transcriptome datasets across many compounds and concentrations.
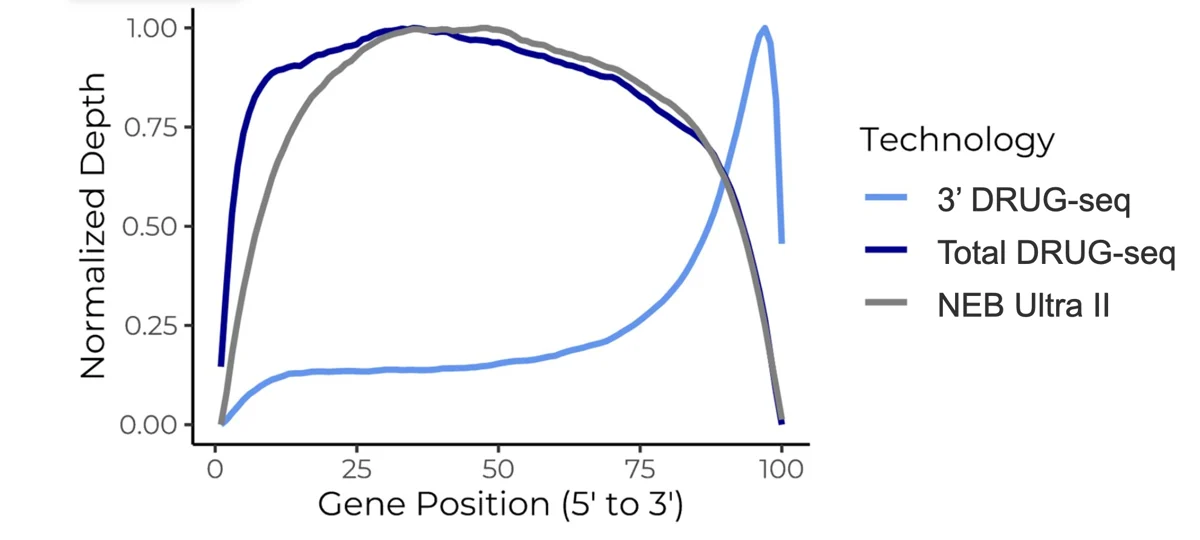
Gene body coverage profiles show the normalized 5’ to 3’ read density across transcript bodies for each library type in Huh7 cells. Total DRUG-seq displays uniform coverage across the full transcript length, matching NEBNext® Ultra™ II Total RNA preparations. Standard DRUG-seq shows characteristic 3’ enrichment, reflecting its 3’-end capture design.
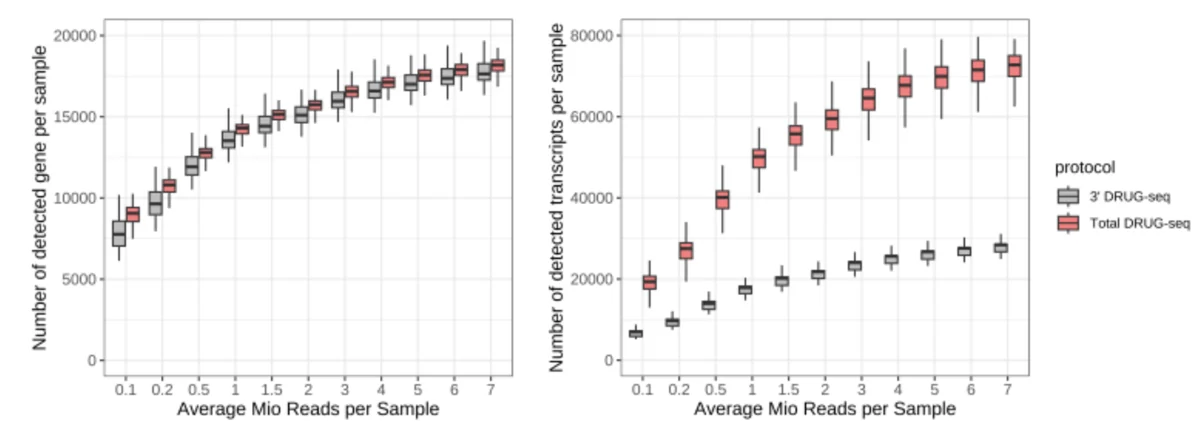
Gene-level (left) and transcript-level (right) detection as a function of sequencing depth after downsampling to between 0.1 and 7 million reads per sample. Both protocols detect comparable numbers of genes across all read depths. At the transcript level, Total DRUG-seq detects up to ~2.5-fold more transcripts than 3′ DRUG-seq at equivalent depths, reflecting its full-length coverage and ability to distinguish isoforms.
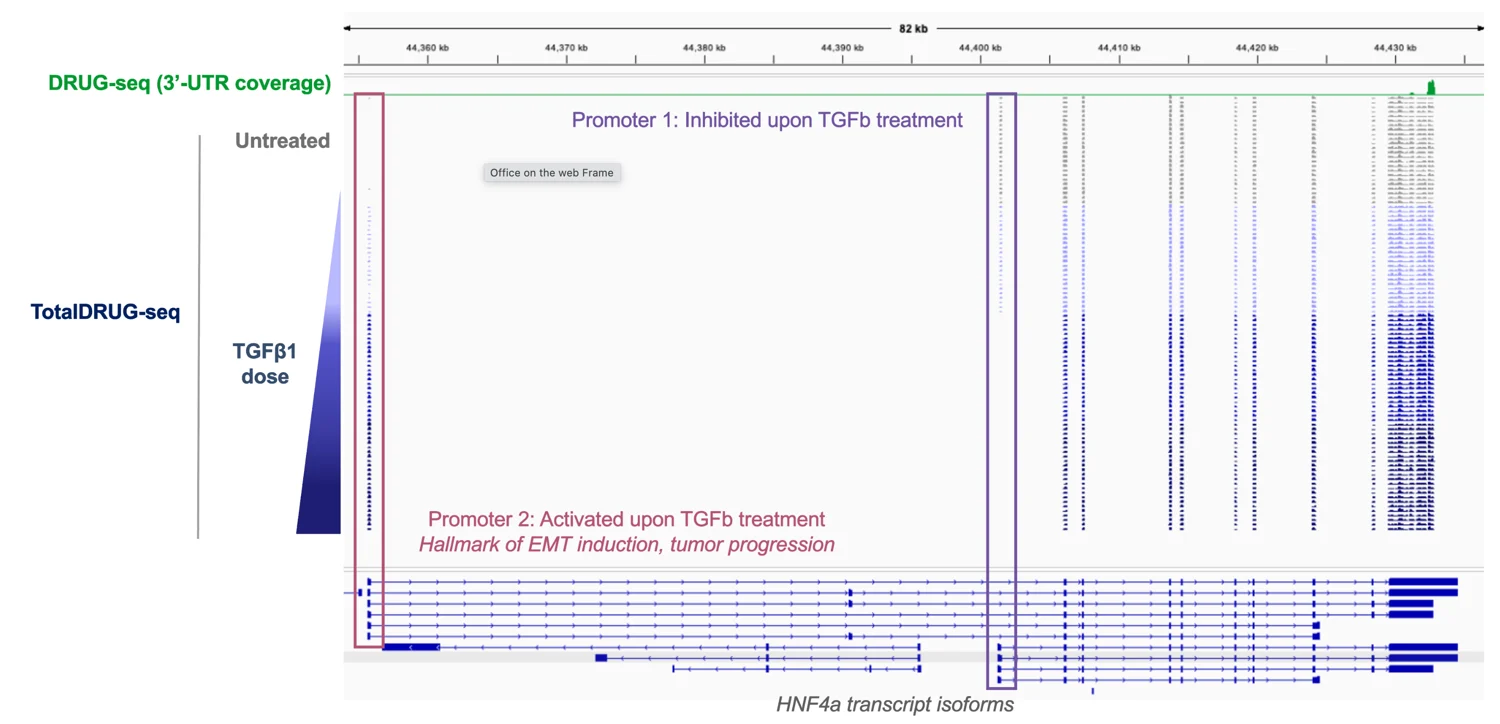
The HNF4α locus has differential isoform usage in response to increasing TGFβ1 dose. Total DRUG-seq (blue tracks) reveals a dose-dependent shift from Promoter 1-driven isoforms (blue box), which are progressively silenced upon TGFβ1 treatment, to Promoter 2-driven isoforms (red box), a hallmark of epithelial-to-mesenchymal transition (EMT) and tumor progression. This demonstrates Total DRUG-seq’s ability to detect biologically meaningful isoform switching events in drug response studies. 3′ DRUG-seq (green track) captures only the shared 3′-UTR region.
| HepaRG | Liver |
| HepG2 | Liver (HCC) |
| Huh7 | Liver (HCC) |
| Hep3B | Liver (HCC) |
| PHH (Primary Human Hepatocytes) | Primary liver |
| NCI-H295R | Adrenal/liver metabolism |
| MCF7 | Breast cancer |
| A549 | Lung carcinoma |
| H358 | Lung cancer |
| NCI-H1563 / H1048 | Lung cancer |
| DLD-1 | Colorectal cancer |
| SW837 | Colorectal cancer |
| HCT116 | Colorectal cancer |
| LS180 | Colorectal cancer |
| COLO201 | Colorectal cancer |
| C2BBe / C2BBe1 | Colorectal |
| GP5d | Colorectal |
| U2OS | Osteosarcoma |
| A172 | Glioblastoma |
| PSN-1 | Pancreatic cancer |
| AsPC-1 | Pancreatic cancer |
| SU.86.86 | Pancreatic cancer |
| A375 | Melanoma |
| HaCaT | Keratinocyte |
| UMUC3 | Bladder cancer |
| 5637 | Bladder cancer |
| HT1197 | Bladder cancer |
| Cal29 | Bladder cancer |
| UBLC1 | Bladder cancer |
| PBMC | Primary blood |
| Jurkat | T-cell leukemia |
| Raji | B-cell lymphoma |
| THP-1 | Monocyte |
| U937 | Monocyte |
| MV-4-11 | AML |
| MOLM13 | AML |
| HL60 | Leukemia |
| MM1S | Myeloma |
| KMS12BM | Myeloma |
| CD4+ / CD8+ T cells | Primary immune |
| Tregs / TILs | Immune subsets |
| CD3 T cells | Immune |
| iPSC | Pluripotent |
| iPSC-derived neurons | Neural |
| iPSC-derived cardiomyocytes | Cardiac |
| iPSC-derived cortical neurons | Neural |
| iPSC-derived organoids | Various |
| HMC3 | Microglia |
| Luhmes | Dopaminergic neuron |
| IMR90 | Fibroblast (used in brain spheres) |
| iCell GlutaNeurons | Neurons |
| Astrocytes (human/mouse/rat) | Glial |
| NHDF | Dermal fibroblast |
| MEF | Mouse embryonic fibroblasts |
| HEK293 / HEK293T | Kidney (transformed) |
| ARPE-19 | Retinal epithelium |
| NHEK | Keratinocytes |
| Lung epithelial | Epithelial |
| RPTEC/TERT1 | Kidney (proximal tubule) |
| T84 | Colon epithelium |
| Human adipocytes | Primary |
| Brown adipocytes | Metabolic |
| Visceral adipocytes | Primary |
| Mouse/canine adipose | Animal |
Each Total DRUG-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different BRB-seq pools.
To note, the total number of RNA samples that can be processed with one kit does not exceed the kit specifications; for instance, a 96-sample kit can be used to prepare up-to 96 samples distributed across up to four different libraries.
The recommended range of input material is in the range of 5’000-50’000 cells.
The only difference between Total DRUG-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the DRUG-seq barcode sequence.
For a thorough description of Total DRUG-seq data processing, please refer to the Total DRUG-seq kit user guide.
The barcode set for your kit is conveniently located on the kit label. Please refer to the label for accurate identification.
For optimal compatibility, ensure that you use the appropriate plate format (e.g., for kits designed for 96 reactions, the 96 well-plate format should be used). This ensures accurate and efficient processing of your samples. If you have any further questions or concerns, please contact our support team for assistance by email or using our live chat tool.
* Contact us to inquire about compatibility with other species.
Product
Number of Samples
MERCURIUS™ Cell lysis module
MERCURIUS™ Cell lysis module
MERCURIUS™ UDI X-Leap Expansion module
MERCURIUS™ Full-Length Post-Pooling Preparation Module (4 libraries)
Each kit includes 4 UDI pairs. Add the expansion module if you need more unique indexes (total 16 UDI pairs available).
Biomarker discovery using gene expression signatures has transformed pharmaceutical discovery and development, from early hit triage and target validation to toxicology, dose optimization, and patient…
Next-generation AI models for drug discovery are only as powerful as the quality, size, and perturbation-richness of their training data. Yet no publicly available dataset…
As the SOT Annual Meeting and ToxExpo 2026 approaches, we take a pre-conference look at the program through a transcriptomics-focused lens. One overall shift is…