Unlock the power of unbiased transcriptome profiling with massively multiplexed, ultra-high-content, and high-throughput RNA sequencing.
MERCURIUS™ BRB-seq turns purified RNA into sequencing-ready libraries in a workflow built for large studies. By barcoding samples early and pooling them upstream, BRB-seq reduces hands-on time and per-sample cost while delivering robust gene expression data across hundreds of samples in parallel. It is a practical fit for cohorts, time-courses, screening-style designs, and any study where traditional sample-by-sample library prep becomes too slow or too expensive.
Early sample barcoding and pooling make it possible to process up to 384 samples together, helping you run larger studies more efficiently.
Choose 3′, Full-Length, or Total RNA readouts depending on whether your priority is scale, isoforms and splicing, or broader whole-transcriptome coverage.
A one-day workflow and low per-sample cost make BRB-seq a strong fit for cohorts, toxicology, plant studies, and large experimental designs.
MERCURIUS™ BRB-seq enables high-throughput transcriptomic profiling across hundreds of samples in parallel, supporting detection of subtle, dose-dependent gene expression changes after toxicant exposure. Its scalability and sensitivity accelerate toxicity biomarker discovery for safer, more efficient drug and chemical development.
MERCURIUS™ BRB-seq offers a scalable, cost-effective solution for transcriptomic analysis in large cohorts and population studies. An integrated globin mRNA depletion step improves transcriptome coverage and data quality from whole blood samples. The technology also performs well with low-input or partially degraded RNA, helping unlock archived blood samples for precision medicine, epidemiology, and disease research.
MERCURIUS™ BRB-seq enables rapid, high-throughput transcriptome analysis at a fraction of the cost of traditional RNA-seq, making it easy to study gene expression across large sample sets. It is well-suited for uncovering gene function, dissecting regulatory networks, and exploring development, differentiation, and cellular response mechanisms in both basic and systems-level research.
By enabling transcriptome profiling across hundreds of samples at once, MERCURIUS™ BRB-seq accelerates discovery of genes linked to agronomic traits such as yield, drought tolerance, disease resistance, and nutrient use efficiency. It helps researchers decode plant responses to biotic and abiotic stress, identify key regulatory pathways, and study plant-pathogen interactions, even in crops with large, complex genomes like wheat and maize.
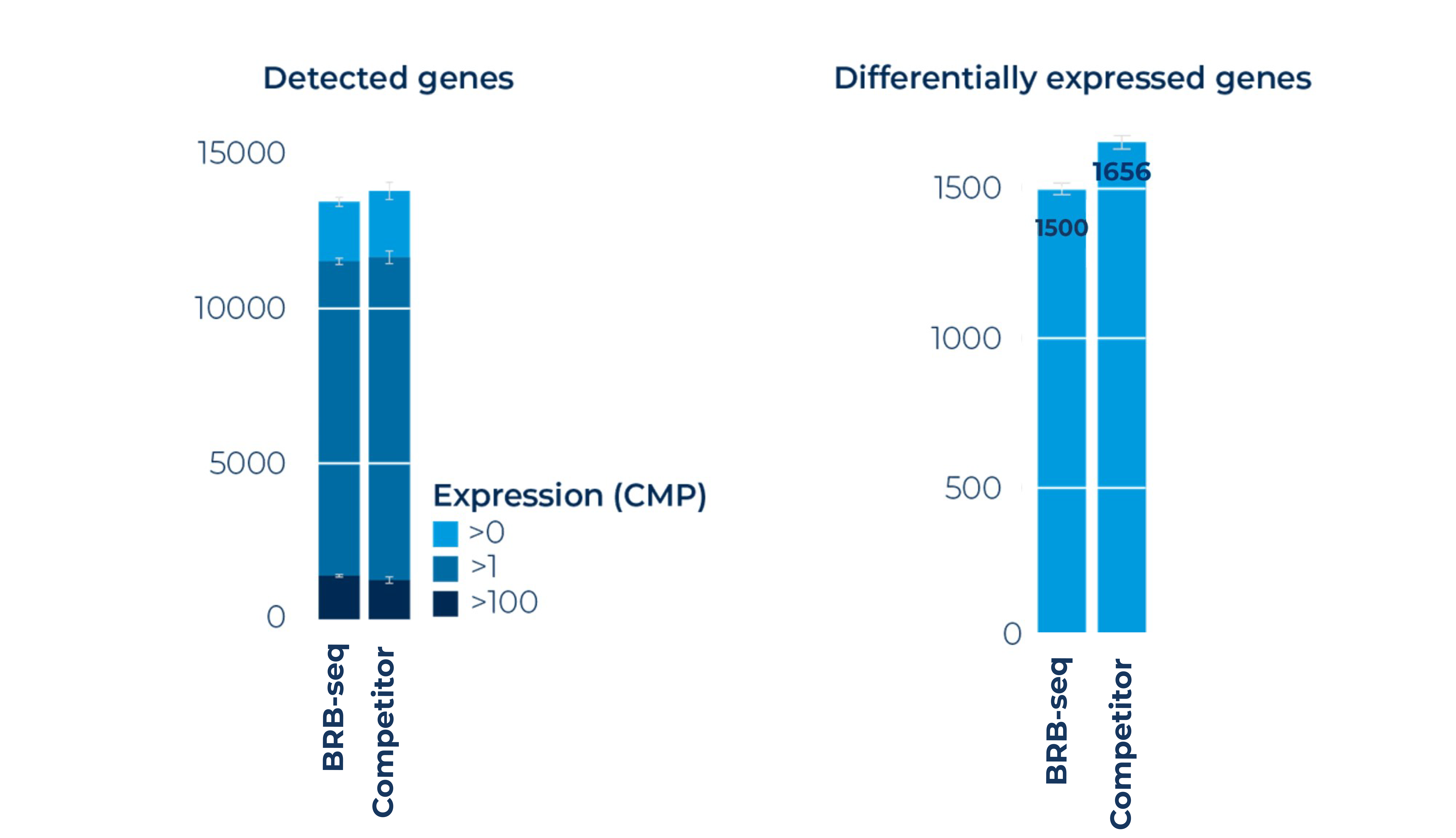
Comparison of gene detection (left) and differential expression (right) between the same samples prepared with MERCURIUS™ BRB-seq or Illumina TruSeq. (Left) Both protocols detect equivalent numbers of genes across all expression thresholds at the same sequencing depth. (Right) Differential expression analysis yields comparable numbers of differentially expressed genes for both technologies. MERCURIUS™ BRB-seq achieves TruSeq-equivalent analytical performance at significantly higher throughput and lower cost per sample.
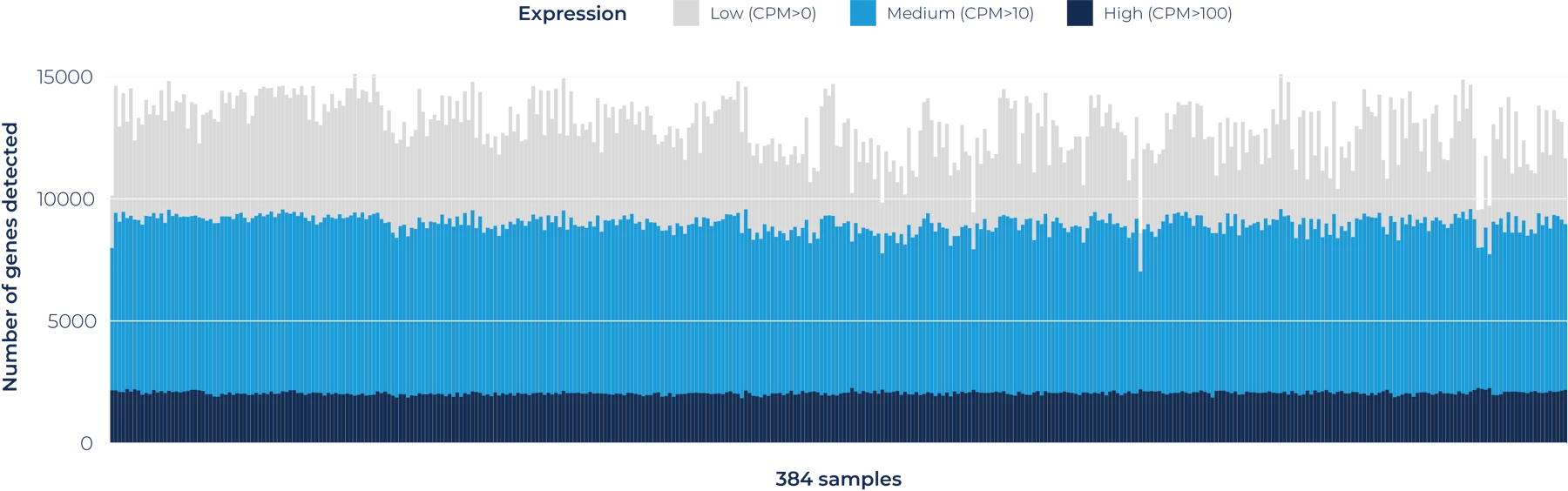
Number of genes detected per sample across a 384-sample BRB-seq run, stratified by expression level. Across all 384 samples, an average of ~13,000 genes is detected when sequenced at a depth of 1.5M reads/sample. The uniformity of gene detection across all samples demonstrates the reproducibility and scalability of MERCURIUS™ BRB-seq for high-throughput transcriptomic profiling.
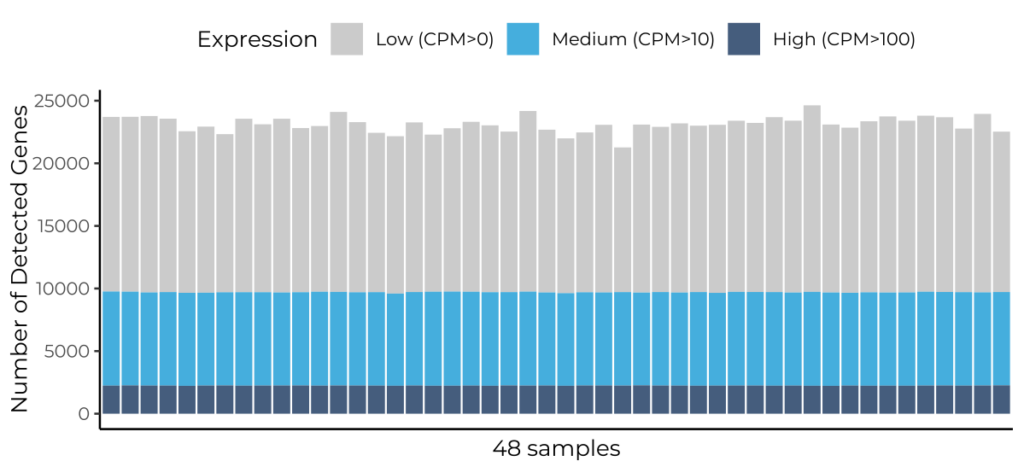
Number of genes detected per sample across a 48-sample MERCURIUS™ Full-Length BRB-seq run, stratified by expression level. All 48 samples show highly uniform gene detection, with approximately 22,000–24,000 genes detected at 12M reads/sample. The consistency across samples demonstrates the reproducibility of MERCURIUS™ Full-Length BRB-seq for high-throughput transcriptomic profiling.
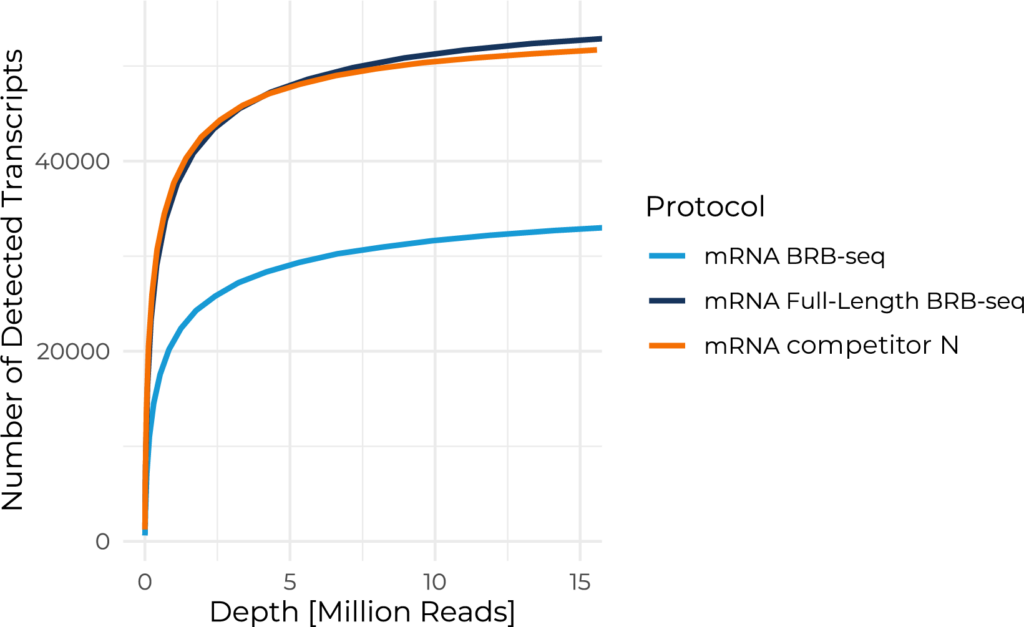
Number of detected transcripts as a function of downsampled sequencing depth for different technologies. MERCURIUS™ Full-Length BRB-seq closely tracks Competitor N across all read depths. Standard BRB-seq detects fewer transcripts, reflecting its 3′-capture design, which limits isoform resolution. These results demonstrate that MERCURIUS™ Full Length BRB-seq delivers competitor-equivalent transcript detection at higher throughput and lower cost.
Saturation curves indicating the number of detected genes as a function of sequencing depth in Huh7 cells. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq perform comparably to NEBNext® Ultra™ II mRNA and Total RNA preparations, respectively, demonstrating equivalent sensitivity at a fraction of the library preparation cost and effort. MERCURIUS™ BRB-seq is a 3’ mRNA library preparation technology, hence the lower overall gene detection versus full-length methods.
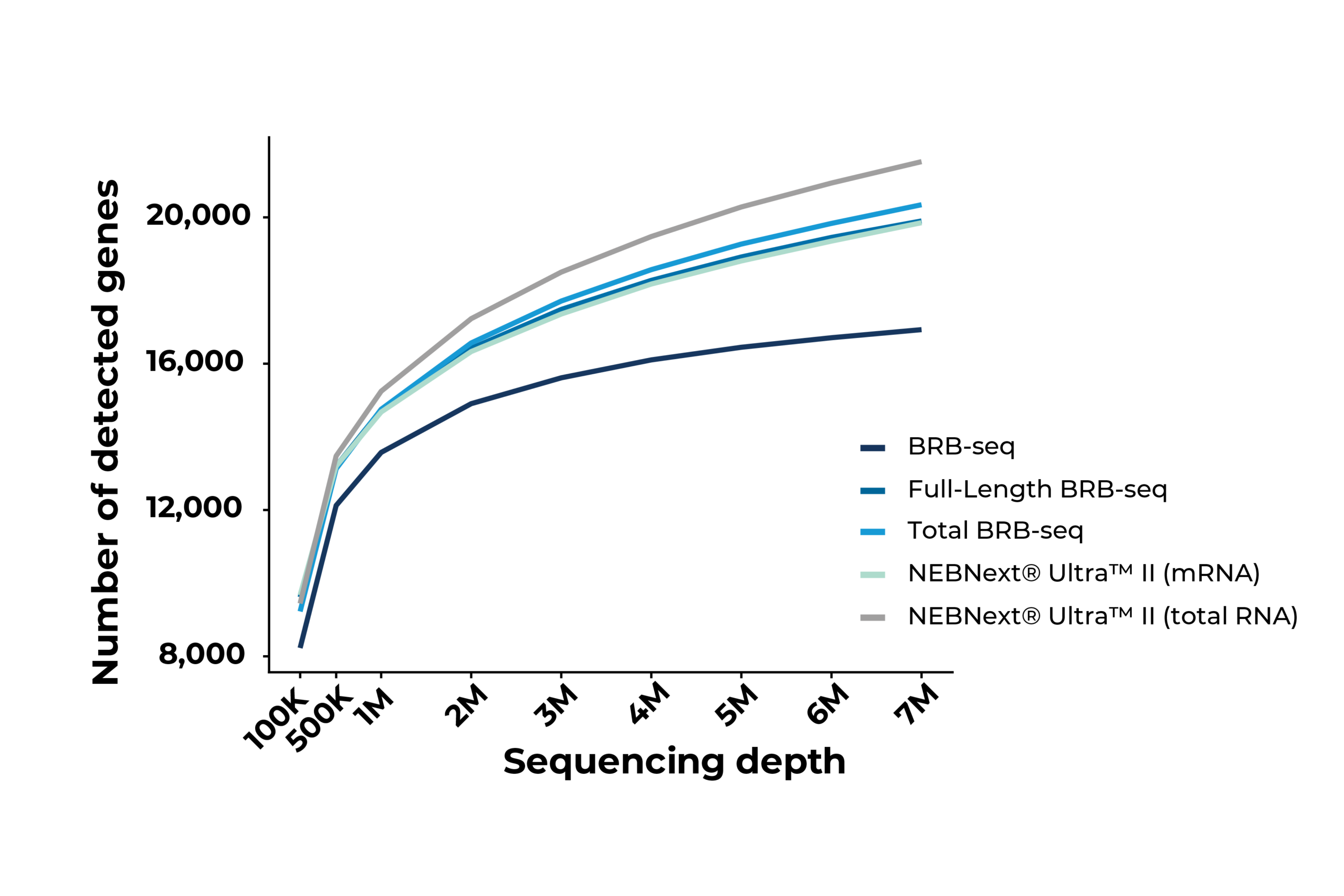

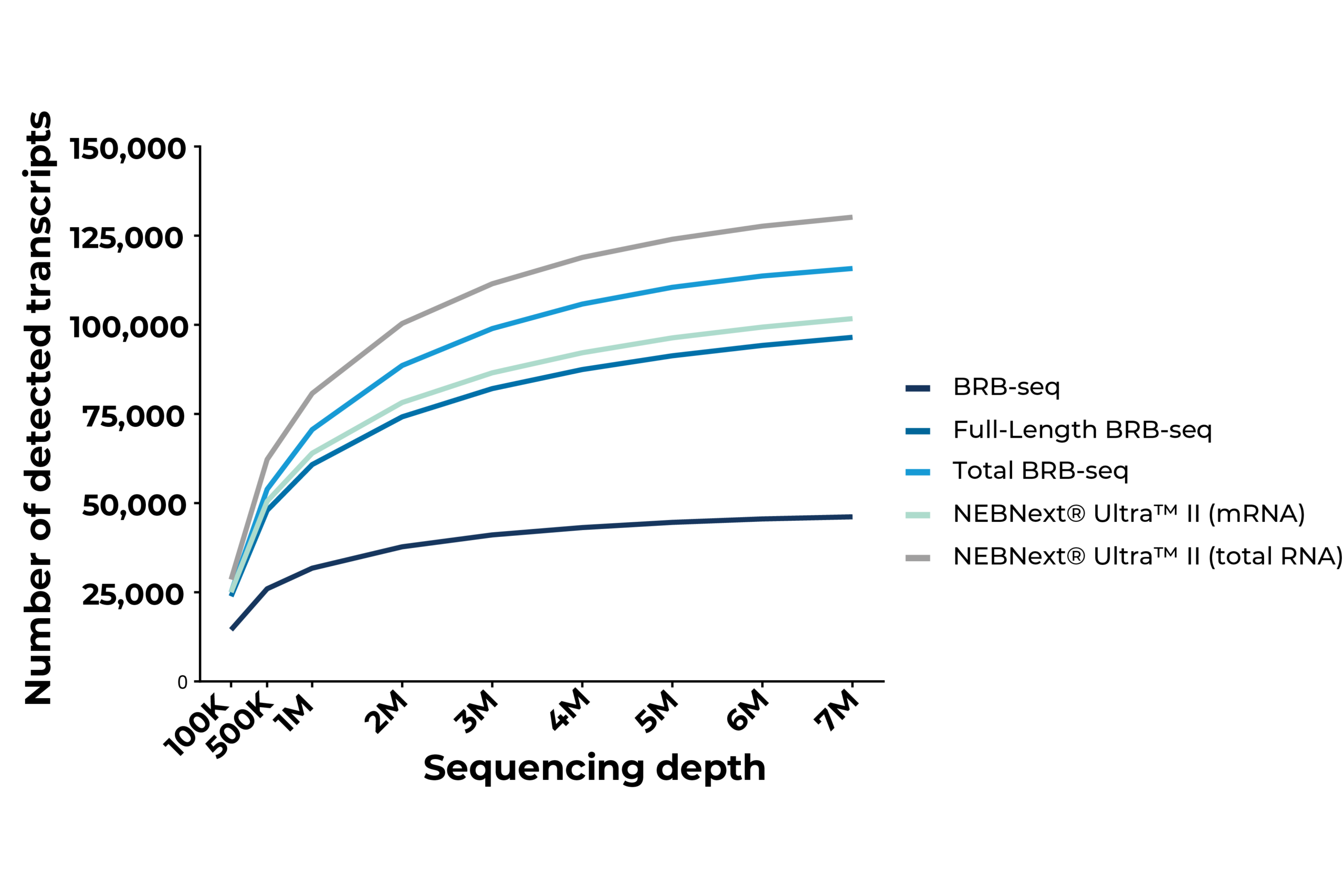
Saturation curves indicating the number of detected transcripts as a function of sequencing depth in Huh7 cells. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq perform comparably to NEBNext® Ultra™ II mRNA and Total RNA preparations, respectively, demonstrating equivalent sensitivity at a fraction of the library preparation cost and effort. MERCURIUS™ BRB-seq specifically detects protein-coding transcripts, hence the lower overall transcript detection compared to full-length methods.
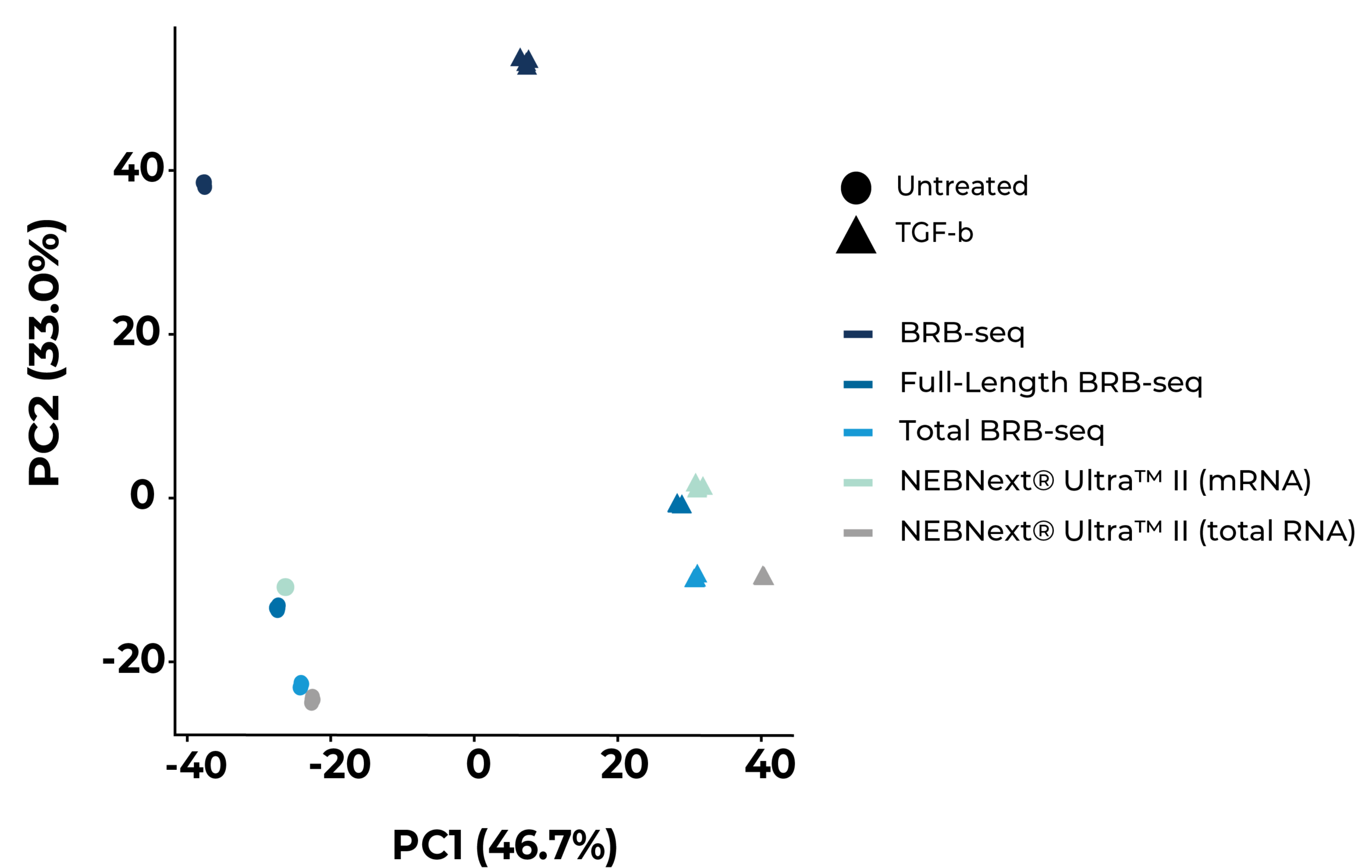
Principal component analysis (PCA) of gene expression profiles generated by the specified library preparation methods across untreated (circles) and TGF-β-treated (triangles) Huh7 samples. Samples cluster primarily by library type along PC2 and by treatment condition along PC1. All methods robustly capture the expression response to treatment. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq cluster closely with their NEBNext® Ultra™ II counterparts, demonstrating equivalent transcriptomic profiles. MERCURIUS™ BRB-seq clusters separately, consistent with its 3′-end capture design.
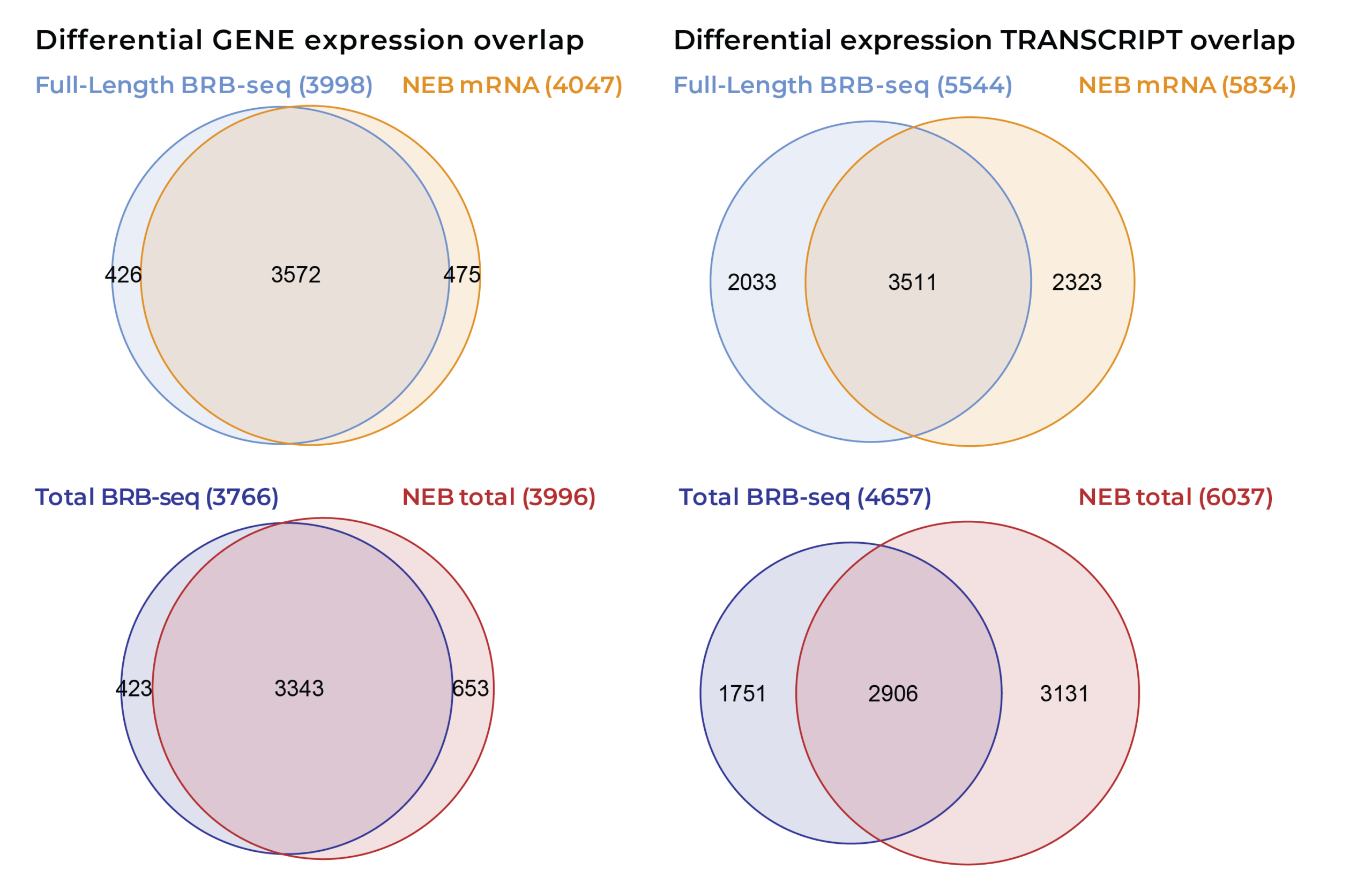
Venn diagrams showing the overlap in differentially expressed (DE) genes (left) and transcripts (right) identified by MERCURIUS™ Full Length BRB-seq and NEBNext® Ultra™ II for mRNA libraries (top) and total RNA libraries (bottom). Numbers indicate DE features unique to each method or shared between both. At the gene level, MERCURIUS™ Full Length BRB-seq and NEBNext® Ultra™ II share over 88% of DE genes for mRNA and 84% for total RNA preparations, demonstrating strong concordance in differential expression calls across both library types. The overlap is smaller but still strong for DE transcripts across technologies.
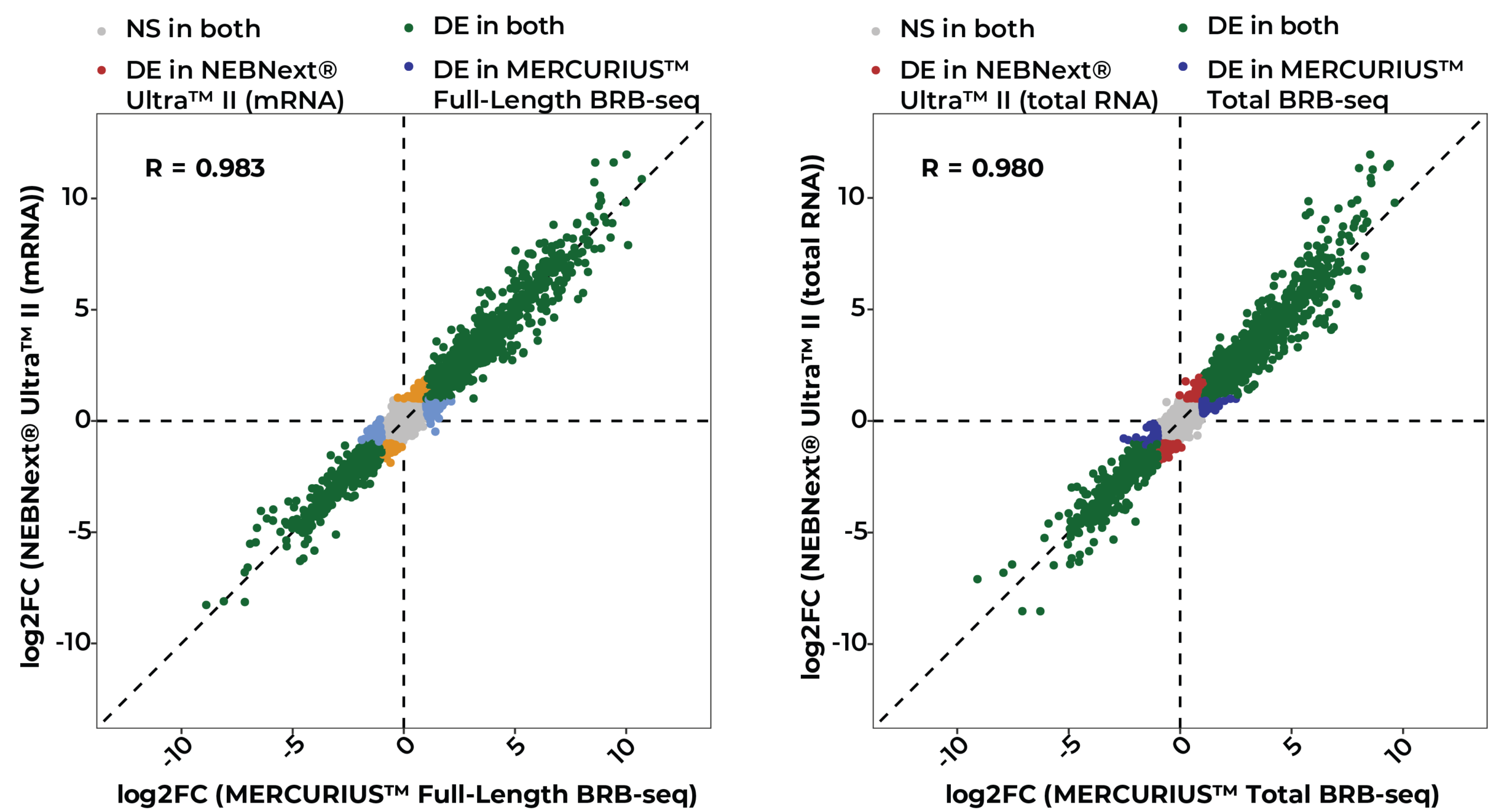
Log2 fold-change correlation between MERCURIUS™ Full-Length BRB-seq and NEBNext® Ultra™ II for mRNA (left, R = 0.983) and total RNA (right, R = 0.980) protocols, across intersecting gene sets of 13,364 and 13,237 genes, respectively. Each point represents a gene, colored by its differential expression (DE) or non-significant (NS) status after treatment of Huh7 cells with human TGF-β versus untreated cells. The dashed diagonal represents perfect correlation. The concordance of DE fold changes across both RNA input types demonstrates that MERCURIUS™ Full-Length and Total BRB-seq reliably recapitulate the results of gold-standard library preparation.
BRB-seq
• 3' mRNA sequencing
• Early multiplexing of up to 384 samples
• 13,000 genes detected at 1.5M reads/sample
• Ideal for comprehensive genome-wide gene expression profiling
Full-Length BRB-seq
• Full-length mRNA sequencing
• Early multiplexing up to 96 samples
• 22,000+ genes detected at 12M reads/sample
Total BRB-seq
• Full-length, total RNA sequencing
• Early multiplexing up to 384 samples
• 25,000+ genes detected at 10M reads/sample
High-Sensitivity BRB-seq
• 3' mRNA sequencing
• Early multiplexing of up to 384 samples
• For low input RNA samples: as low as 1ng/sample
• 15,000+ genes detected at 5M reads/sample

BRB-seq service
By leveraging BRB-seq, we not only provide industrial and academic clients with quality RNA-seq data, but we also do so with the highest affordability and quick turnaround times.

High-Sensitivity BRB-seq service
This service offers a convenient and streamlined solution for transcriptomics projects with a limited amount of input RNA (starting from 1 ng per sample).

Full-Length BRB-seq service
Scalable full-length mRNA sequencing for purified RNA, combining early multiplexing with isoform-level, splicing, and transcript-structure insights.

Total BRB-seq service
The MERCURIUS™ Total BRB-seq service offers a convenient and streamlined solution for full-length total RNA transcriptomics projects of any size.

Blood BRB-seq service
High-throughput 3′ mRNA-seq service for human whole-blood samples, with integrated globin depletion and expertise in high-throughput RNA extraction from PaxGene tubes.

Total Blood BRB-seq service
Multiplexed total RNA-seq service for human whole-blood samples, with inline globin depletion and high-throughput RNA extraction from PaxGene tubes.

Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…
In this post, we will see briefly how to perform downstream analysis for RNA-seq data.
…MERCURIUS™ BRB-seq is a transformative tool combining unbiased, ultra-high-content, and high-throughput screening with massively parallel transcriptomics.
This method uses highly optimized and rigorously evaluated sample barcodes and unique molecular identifiers to tag the 3’ poly(A) tail of all mRNA molecules in a sample-specific manner during the first-strand synthesis step of cDNA library preparation. After this step, all samples can be pooled into one single tube and processed together for the remainder of the library prep workflow.
The BRB-seq protocol is designed to work with purified RNA samples from all eukaryotic species.
MERCURIUS™ BRB-seq stands out from traditional RNA-seq methods by offering a high-throughput, cost-efficient, and streamlined workflow specifically designed for large-scale screening applications. One of the key differences lies in its sample multiplexing strategy: BRB-seq allows multiple RNA samples to be barcoded and pooled at the earliest step of the protocol—right after the RT reaction—so that the entire library preparation can proceed in a single tube. This significantly reduces both hands-on time and reagent costs.
In contrast, standard RNA-seq workflows typically require individual processing of all the RNA samples. This makes them more labor-intensive, costly, and less scalable, especially when working with large numbers of conditions, compounds, or replicates—common in drug discovery pipelines.
Each BRB-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different BRB-seq pools. To note, the total number of RNA samples that can be processed with one kit does not exceed the kit specifications; for instance, a 96-samples kit can be used to prepare up-to 96 samples distributed across up-to four different libraries.
One of the key advantages of BRB-seq is that it does not only save reagents and cost in the library preparation stage, but also in the sequencing one. As opposed to standard RNA-seq, where 20M-30M reads per sample are required, we normally recommend to sequence BRB-seq libraries at a depth of 4M-5M reads per sample, which is normally enough to detect the vast majority of expressed genes.
The only difference between BRB-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the BRB-seq barcode sequence. For a thorough description of BRB-seq data processing, please refer to the BRB-seq kit user guide.
Tell us about your project and we will help you find the right approach.