MERCURIUS™ Total Blood BRB-seq is a library preparation kit for purified human blood RNA that combines early multiplexing, inline globin depletion, and full-length total RNA sequencing in one workflow. It is designed to generate sequencing-ready libraries for both coding and non-coding RNAs, while also supporting analyses such as transcript variants, alternative splicing, and fusion genes. The kit is available in 96- and 384-prep formats, multiplexes 96 samples in a single tube, includes globin blockers, and is compatible with Illumina® and AVITI™ sequencing platforms.

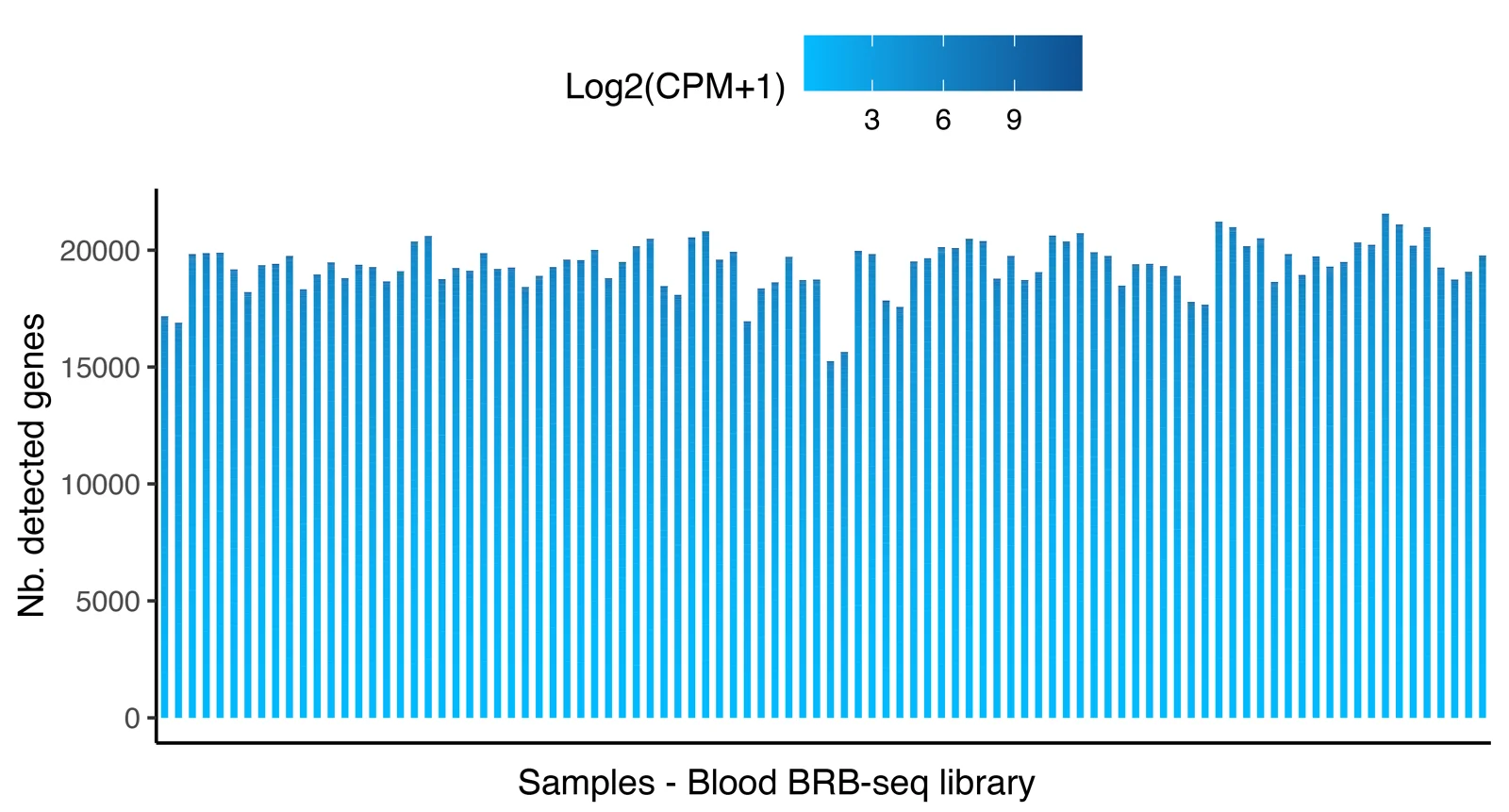
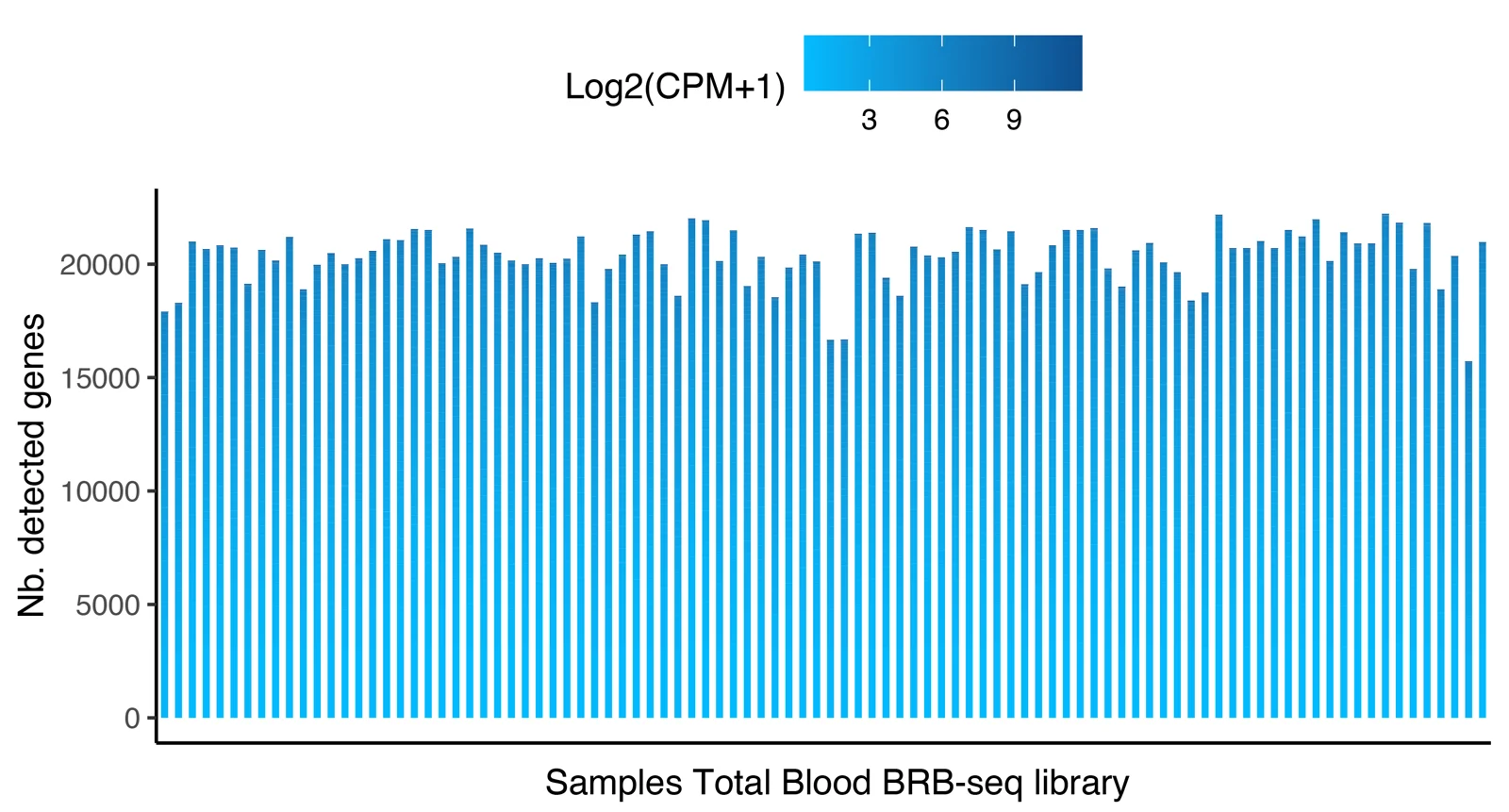
Distribution of the number of detected genes across 96 samples prepared with the MERCURIUS™ Blood BRB-seq library preparation kit and the MERCURIUS™ Total Blood BRB-seq. The libraries were sequenced at the same sequencing depth.
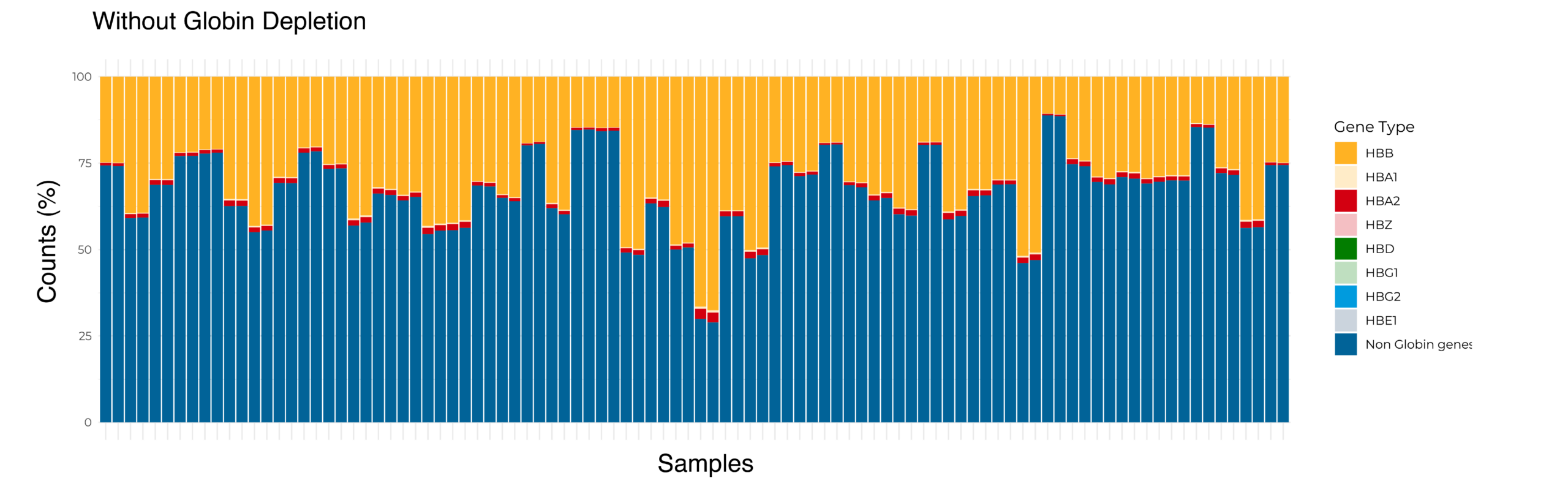
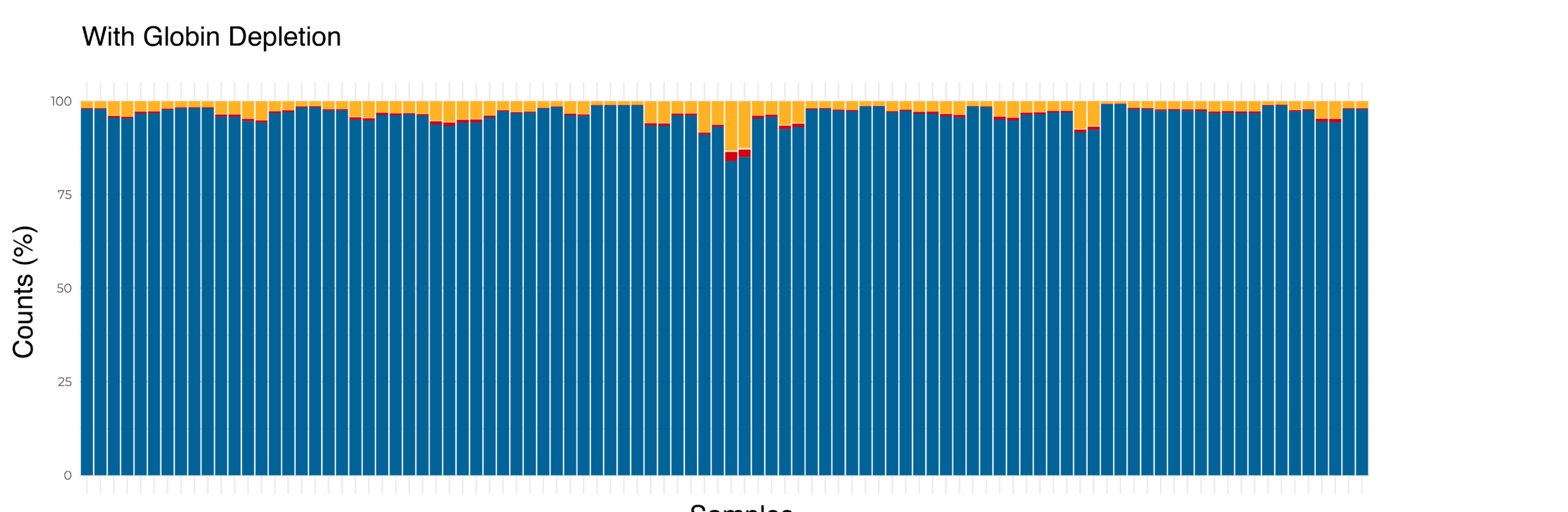
(Top) The distribution of reads mapped to non-globin and globin genes per sample for one 96 sample Total Blood BRB-seq library without globin depletion. An average of approximately one-third of reads mapped to globin genes. (Bottom) The distribution of reads mapped to non-globin and globin genes per sample for the same 96 samples in one Total Blood BRB-seq library treated with globin depletion. The reads associated with globin genes are drastically reduced, with more reads mapped to non-globin genes.
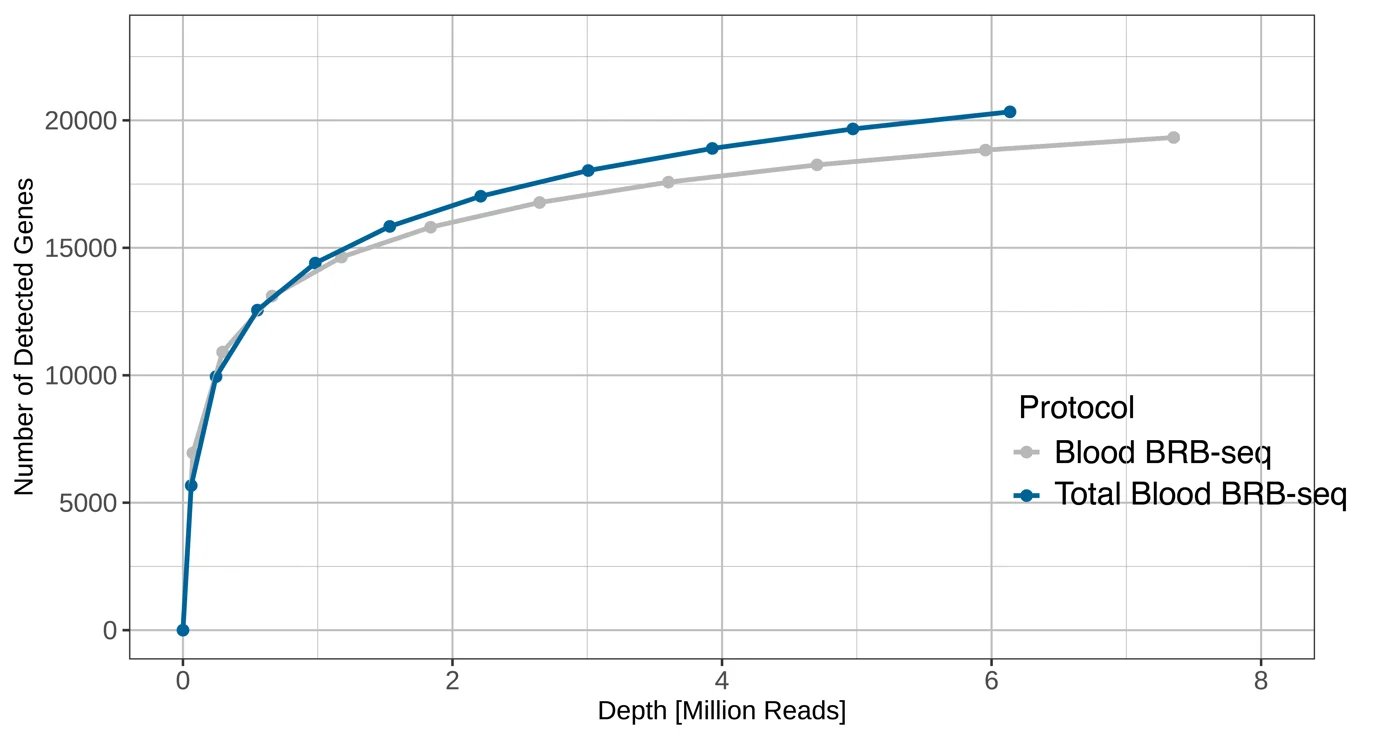
Number of detected genes at different downsampled sequencing depths for Total Blood BRB-seq and Blood BRB-seq. Total Blood BRB-seq consistently detects more genes across the full depth range, reaching approximately 20,000 genes at 6 million reads compared to ~19,000 for Blood BRB-seq.
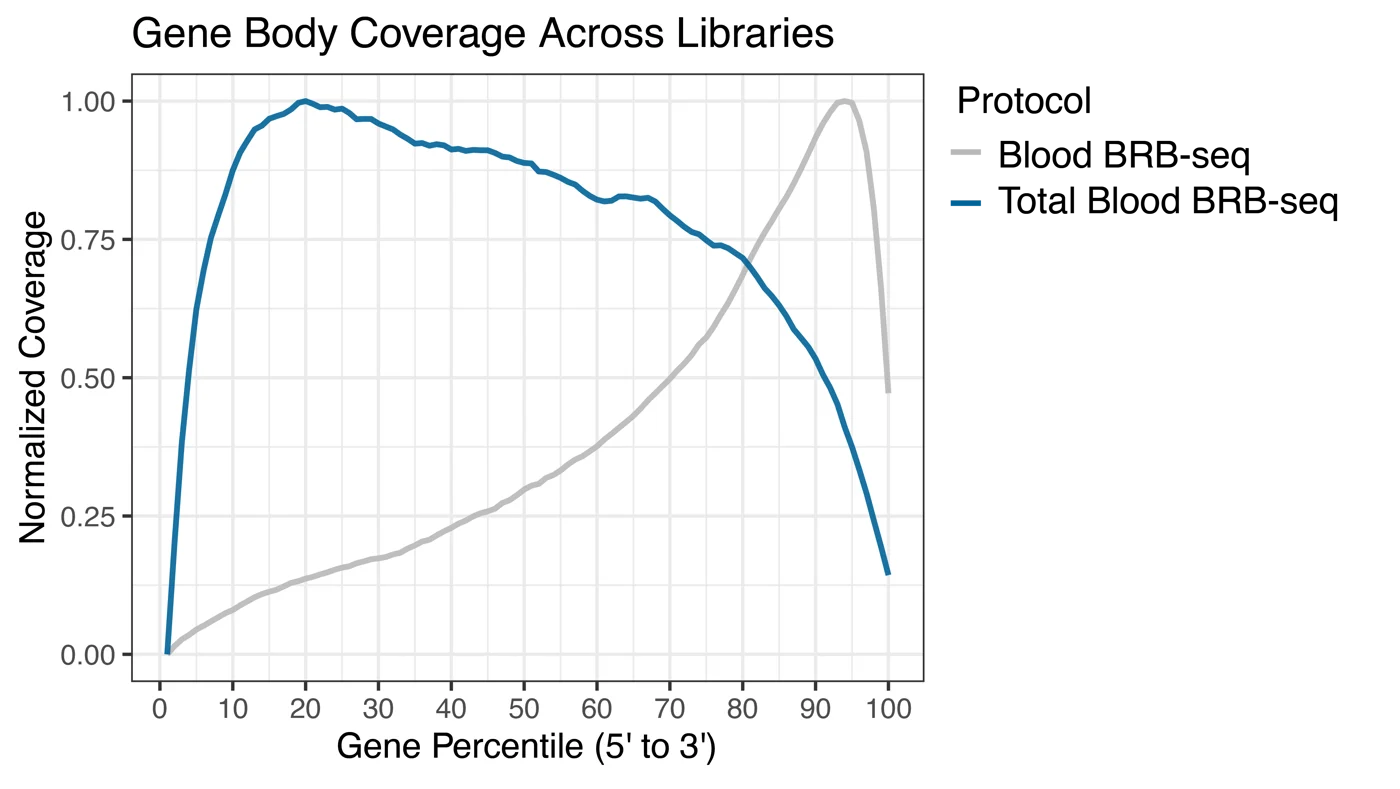
Normalized read coverage across gene transcript bodies for Total Blood BRB-seq and Blood BRB-seq. Total Blood BRB-seq produces a broadly uniform coverage profile across the full transcript length, with coverage peaking early and remaining consistently high through to the 80th gene percentile. Blood BRB-seq displays a pronounced 3′-end bias, owing to its 3′-capture design.
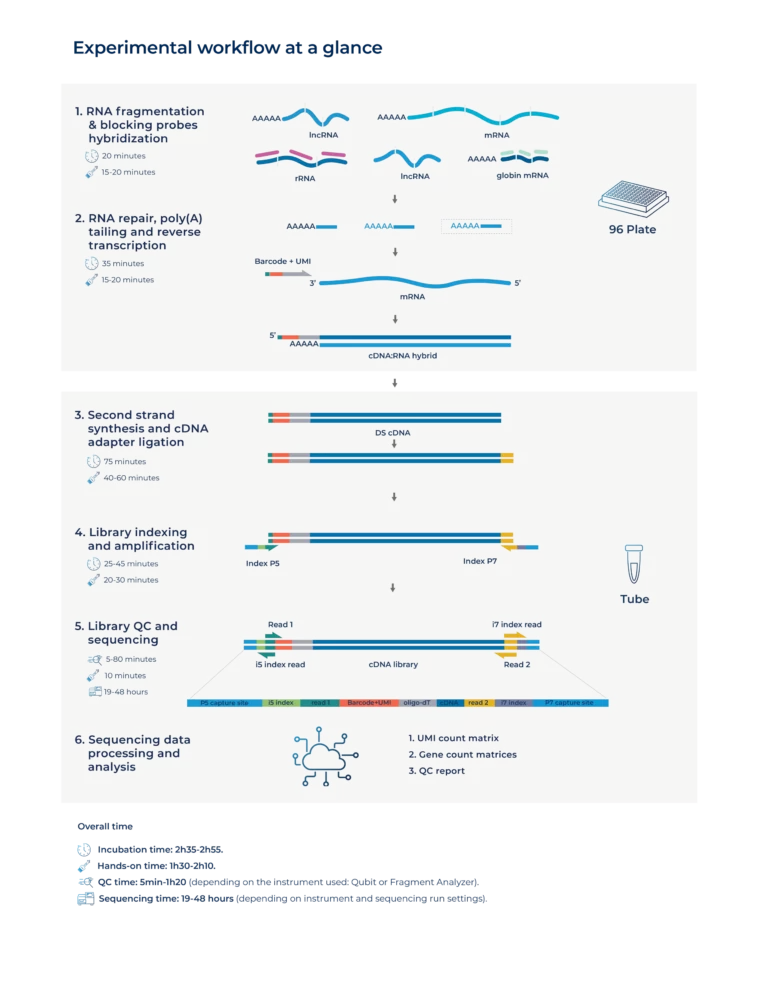
Each Total Blood BRB-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different BRB-seq pools.
To note, the total number of RNA samples that can be processed with one kit does not exceed the kit specifications; for instance, a 96-sample kit can be used to prepare up to 96 samples distributed across up-to four different libraries.
No, the Total Blood BRB-seq workflow is only dedicated to human blood samples.
The recommended range of RNA amount for each sample is of 10ng-100ng.
The minimum recommended RIN number is 7 and the A260/280 ratio (Nanodrop) should be A260/280 >1.5
The only difference betweenTotal Blood BRB-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the barcode sequence.
For a thorough description of Total Blood BRB-seq data processing, please refer to the kit user guide.
The recommended sequencing depth for the Total Blood BRB-seq libraries is between 2-10 M reads per sample, which is typically enough to detect the vast majority of expressed genes.
Go to the “Resources” section of this page and click on the button to download the barcode files for the product you are using.
Yes. If you are not ready to run kits internally, Alithea can run the workflow and deliver FASTQ files, gene count matrices, and analysis report files.
Product
Catalog Number
MERCURIUS™ UDI X-Leap Expansion module
MERCURIUS™ Full-Length Post-Pooling Preparation Module (4 libraries)
Each kit includes 4 UDI pairs. Add the expansion module if you need more unique indexes (total 16 UDI pairs available).
Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…