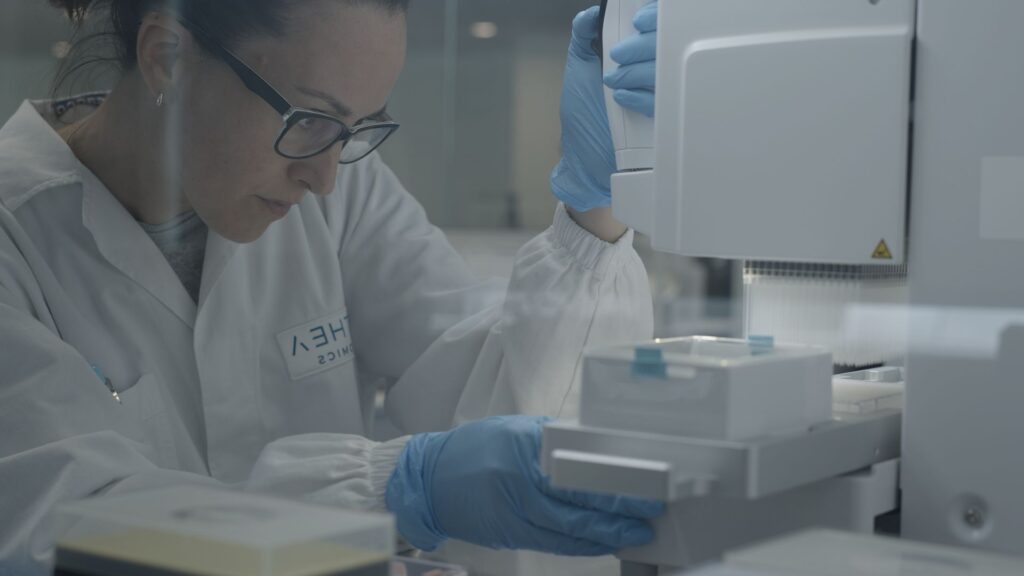
Our DRUG-seq service provides industry and academic scientists with ultra-scalable RNA-seq that enables highly sensitive, RNA-extraction-free, massively multiplexed, and extremely cost-effective transcriptomic screens for large-scale compound discovery programs, next-generation toxicology strategies, and mechanism-of-action studies.
Discover Sample Datasets
HepG2, Hap1, HeLa
The MERCURIUS™ DRUG-seq service is the most comprehensive, high-throughput, and cost-effective solution for large-scale gene expression profiling projects starting from 2D cell cultures.
As part of the DRUG-seq service, users simply need to deliver 96- or 384-well plates with frozen cells to our service centers located in Switzerland and the United States.
Our extensive expertise with DRUG-seq and state-of-the-art sequencing infrastructure allows us to offer one of the most competitive turnaround times and prices on the market.

2 days
(Qubit, Fragment analyzer, shallow sequencing)
1 week
1 week
Raw FASTQ files, sample report file, QC files, and gene count tables
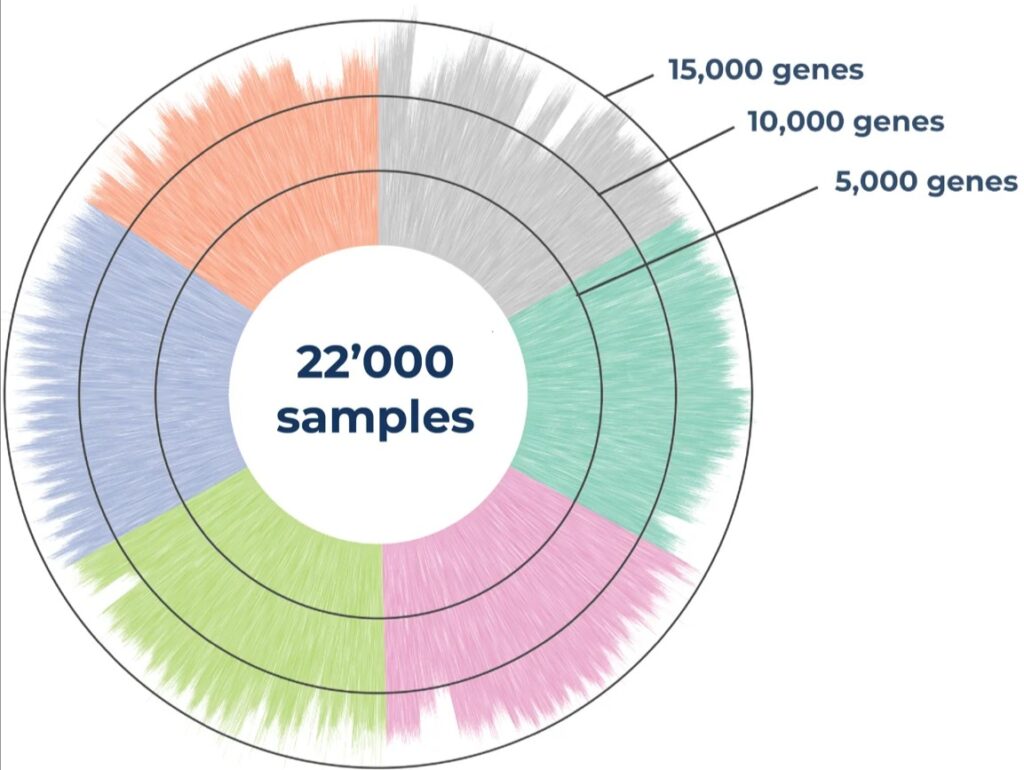
Distribution of the number of detected genes across 22,000 samples in 125x 384-well MERCURIUS™ DRUG-seq plates from six different frozen cell lines. The library was sequenced at an average of 1 million reads per sample on an Illumina NovaSeq 6000.
Compounds
Cell lines
Genes/sample
1M reads/sample
UMAP projection of transcriptional profiles from cells treated with four compounds individually and in combination, alongside untreated controls. Each point represents a single sample, coloured by treatment condition. Samples cluster by transcriptional similarity, revealing that single-compound treatments and combination treatments occupy distinct regions of transcriptional space.
To access the deep-sequenced dataset (7.3 M reads per sample), contact us.
To access the deep-sequenced dataset (3.9 M reads per sample), contact us.
To access the deep-sequenced dataset (7.1 M reads per sample), contact us.
To ensure high-quality data, we typically recommend that each well in a 96-well plate contain 15k-50k cells or 2k-10k cells in a 384-well plate. The minimum number of cells per pool should be 80k.
MERCURIUS™ DRUG-seq can be performed on various sample types, including: cell lines, primary cells, or spheroids (see Spheroid DRUG-seq service).
Check out our DRUG-seq Kits page for a list of our validated cell lines.
DRUG-seq is a 3’-end RNA sequencing method and, as such, requires significantly less sequencing depth than standard full-length RNA-seq to achieve accurate gene quantification. We therefore normally recommend sequencing 1 to 10 million reads per sample, enabling the reliable and unbiased detection of over 18’000 genes.
As part of our standard service pipeline, we align the generated data to the genome of choice, provide a detailed report on the alignment and gene-count statistics, and provide ready-to-use gene-count matrices for downstream analysis.
You can either book a call with our experts to discuss your project or submit your experimental details via our contact form so we can review your design and requirements.
Based on the goals, sample types, and scale of your study, we may recommend starting with a pilot project to optimise conditions and de-risk a larger screen.
If you’re interested in implementing the technology in your own lab instead, you can explore our MERCURIUS™ DRUG-seq kits on the dedicated kits page.
Biomarker discovery using gene expression signatures has transformed pharmaceutical discovery and development, from early hit triage and target validation to toxicology, dose optimization, and patient…
Next-generation AI models for drug discovery are only as powerful as the quality, size, and perturbation-richness of their training data. Yet no publicly available dataset…
As the SOT Annual Meeting and ToxExpo 2026 approaches, we take a pre-conference look at the program through a transcriptomics-focused lens. One overall shift is…