Our Total DRUG-seq service provides industry and academic scientists with ultra-scalable total RNA-seq, enabling whole-transcriptome screens of full-length coding and non-coding transcripts when isoform and alternative splicing detection is crucial. Ideal for large-scale compound discovery programs, next-generation toxicology strategies, and mechanism-of-action studies.
Our MERCURIUS™ Total DRUG-seq service is a convenient and scalable solution for projects of any size, enabling transcriptome-wide detection of full-length coding and non-coding transcripts.
As part of the Total DRUG-seq service, users simply deliver their 96- or 384-well plates containing frozen cells to us in Switzerland or the United States.
Upon receipt of the plates, our team prepares the libraries, then sequences to your desired read depth per sample, and then performs data pre-processing.
We return results, including raw fastq files, sequencing and alignment reports, and gene count matrices suitable for downstream differential expression analyses, once the data meet our rigorous quality control criteria.
During the process, we always keep clients informed at defined checkpoints so we can decide together how best to proceed to the next steps.
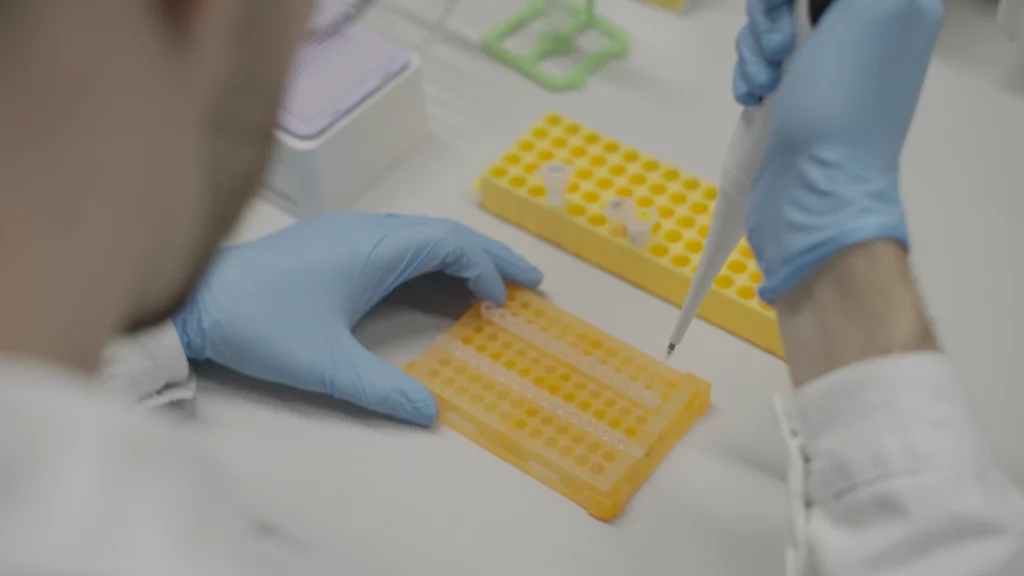

2 days
(Qubit, Fragment analyzer, shallow sequencing)
1 week
1 week
Raw FASTQ files, sample report file, QC files, and gene count tables
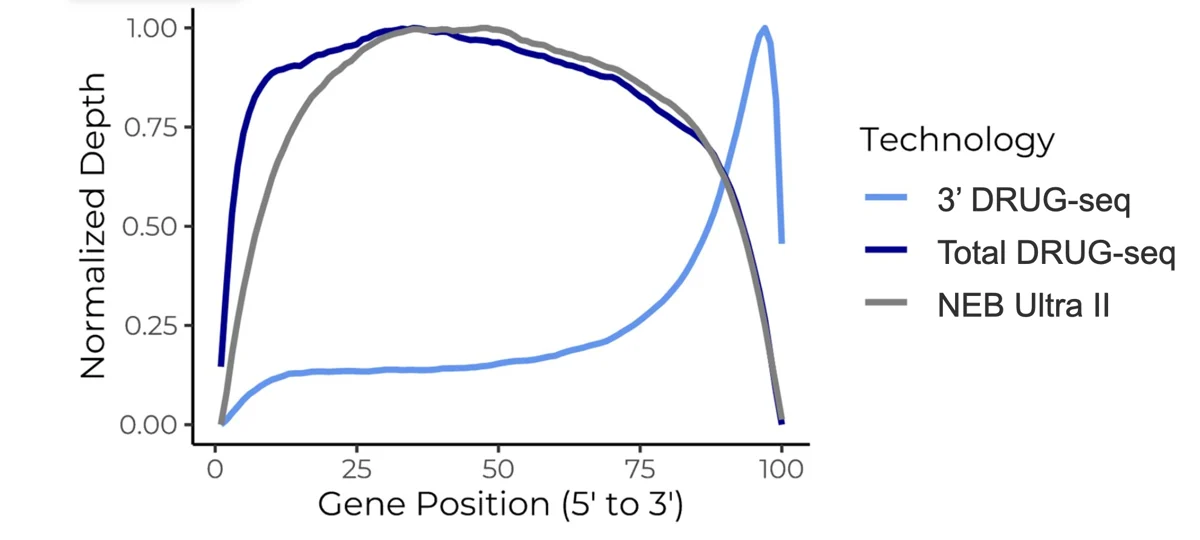
Gene body coverage profiles show the normalized 5’ to 3’ read density across transcript bodies for each library type in Huh7 cells. Total DRUG-seq displays uniform coverage across the full transcript length, matching NEBNext® Ultra™ II Total RNA preparations. Standard DRUG-seq shows characteristic 3’ enrichment, reflecting its 3’-end capture design.
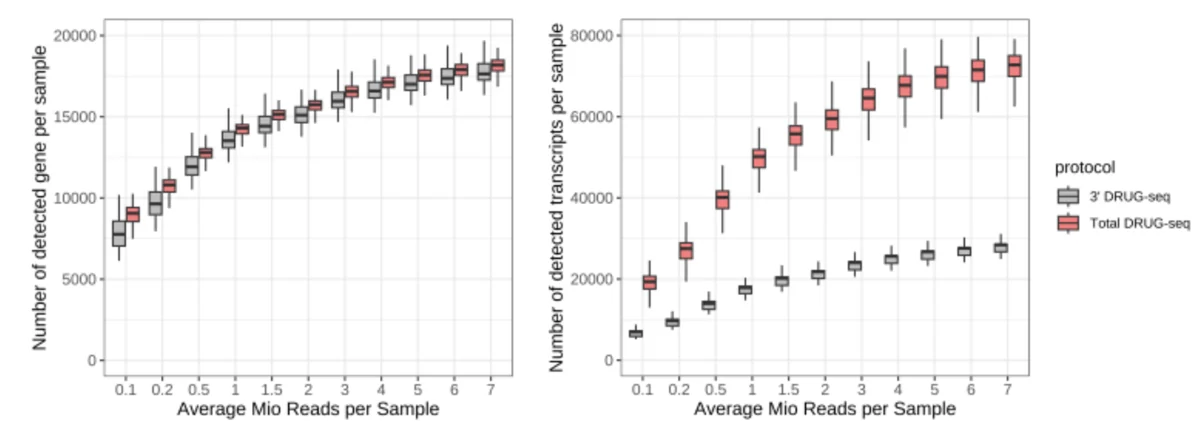
Gene-level (left) and transcript-level (right) detection as a function of sequencing depth after downsampling to between 0.1 and 7 million reads per sample. Both protocols detect comparable numbers of genes across all read depths. At the transcript level, Total DRUG-seq detects up to ~2.5-fold more transcripts than 3′ DRUG-seq at equivalent depths, reflecting its full-length coverage and ability to distinguish isoforms.
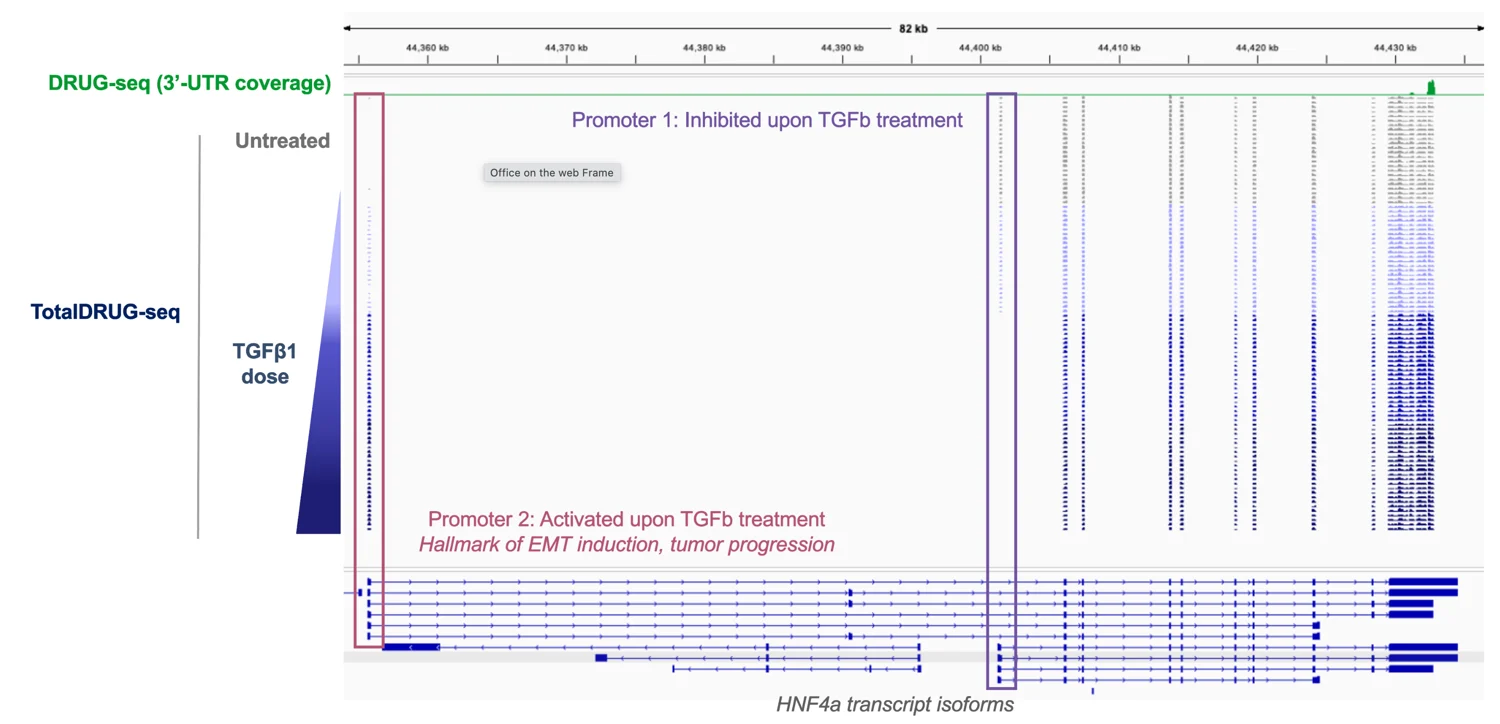
The HNF4α locus has differential isoform usage in response to increasing TGFβ1 dose. Total DRUG-seq (blue tracks) reveals a dose-dependent shift from Promoter 1-driven isoforms (blue box), which are progressively silenced upon TGFβ1 treatment, to Promoter 2-driven isoforms (red box), a hallmark of epithelial-to-mesenchymal transition (EMT) and tumor progression. This demonstrates Total DRUG-seq’s ability to detect biologically meaningful isoform switching events in drug response studies. 3′ DRUG-seq (green track) captures only the shared 3′-UTR region.
To have access to the deep-sequenced dataset (7.3 M reads per sample) contact us.
To have access to the deep-sequenced dataset (7.1 M reads per sample) contact us.
To generate high-quality sequencing data, we recommend starting with 5,000 to 25,000 mammalian cells per well for a 96-well plate and 2,000 to 10,000 cells per well for a 384-well plate.
The total RNA amount per pool should be at least 80,000 cells.
MERCURIUS™ Total DRUG-seq can be performed on various sample types, including: cell lines, primary cells, organoids, or spheroids (see our dedicated service for spheroids).
Check out our Total DRUG-seq Kits page for a list of our validated cell lines.
Total DRUG-seq provides comprehensive coverage of full-length coding and non-coding transcripts.
We therefore recommend sequencing 10 to 20 million reads for each sample, depending on the project’s scale and scope.
You can either book a call with our experts to discuss your project or submit your experimental details via our contact form so we can review your design and requirements.
Based on the goals, sample types, and scale of your study, we may recommend starting with a pilot project to optimise conditions and de-risk a larger screen.
If you’re interested in implementing the technology in your own lab instead, you can explore our MERCURIUS™ Total DRUG-seq kits on the dedicated kits page.
Biomarker discovery using gene expression signatures has transformed pharmaceutical discovery and development, from early hit triage and target validation to toxicology, dose optimization, and patient…
Next-generation AI models for drug discovery are only as powerful as the quality, size, and perturbation-richness of their training data. Yet no publicly available dataset…
As the SOT Annual Meeting and ToxExpo 2026 approaches, we take a pre-conference look at the program through a transcriptomics-focused lens. One overall shift is…