Total Blood BRB-seq enables industrial and academic clients to generate scalable total RNA-seq data directly from whole blood with integrated globin depletion for maximum detection of coding and non-coding transcripts
The MERCURIUS™ Total Blood BRB-seq service offers a convenient and scalable solution for blood transcriptomics projects on human samples that require full-length coding and non-coding transcript detection.
For Total Blood BRB-seq services, customers send purified RNA extracted from eligible human blood samples. Upon receipt, we perform incoming quality control to assess sample suitability before launching the Total Blood BRB-seq pipeline.
Our team then prepares the libraries and sequences to your desired read depth per sample and performs data preprocessing.
Once the data meet our rigorous quality control criteria, we return the results, including raw fastq files, sequencing and alignment reports, and gene count matrices suitable for downstream differential expression analyses.
During the process, we always keep clients informed at defined checkpoints so we can decide together how best to proceed to the next steps.

We accept extracted RNA from human blood samples. See our sample submission guidelines.
Sample submission guidelines
Submit the sample submission form.
(Nanodrop and Fragment analyzer)
1 week
2 days
(Qubit and Fragment analyzer)
1 week
1 week
Raw FASTQ files, sample report file, QC files, and gene count tables
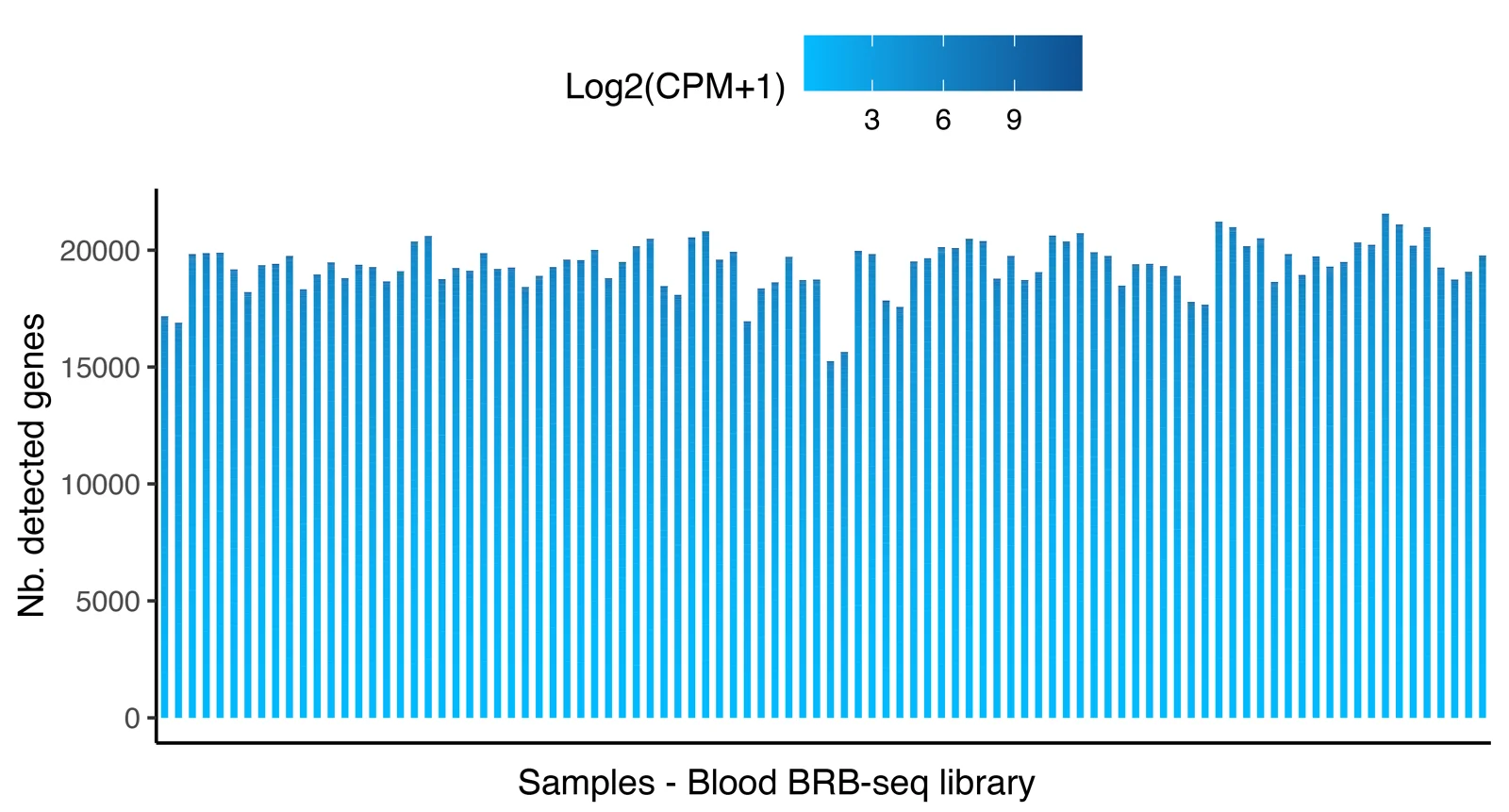
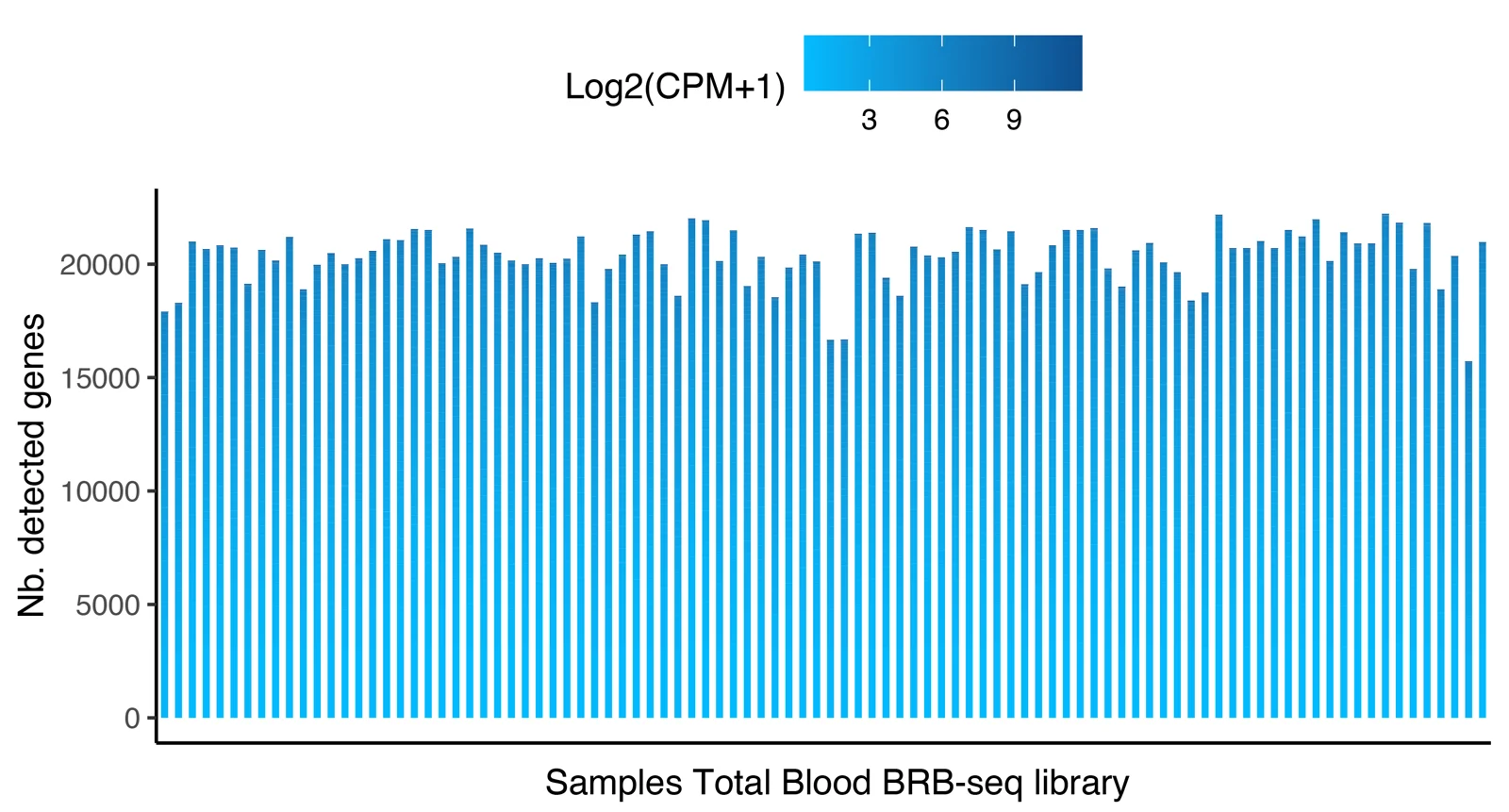
Distribution of the number of detected genes across 96 samples prepared with the MERCURIUS™ Blood BRB-seq library preparation kit and the MERCURIUS™ Total Blood BRB-seq. The libraries were sequenced at the same sequencing depth.
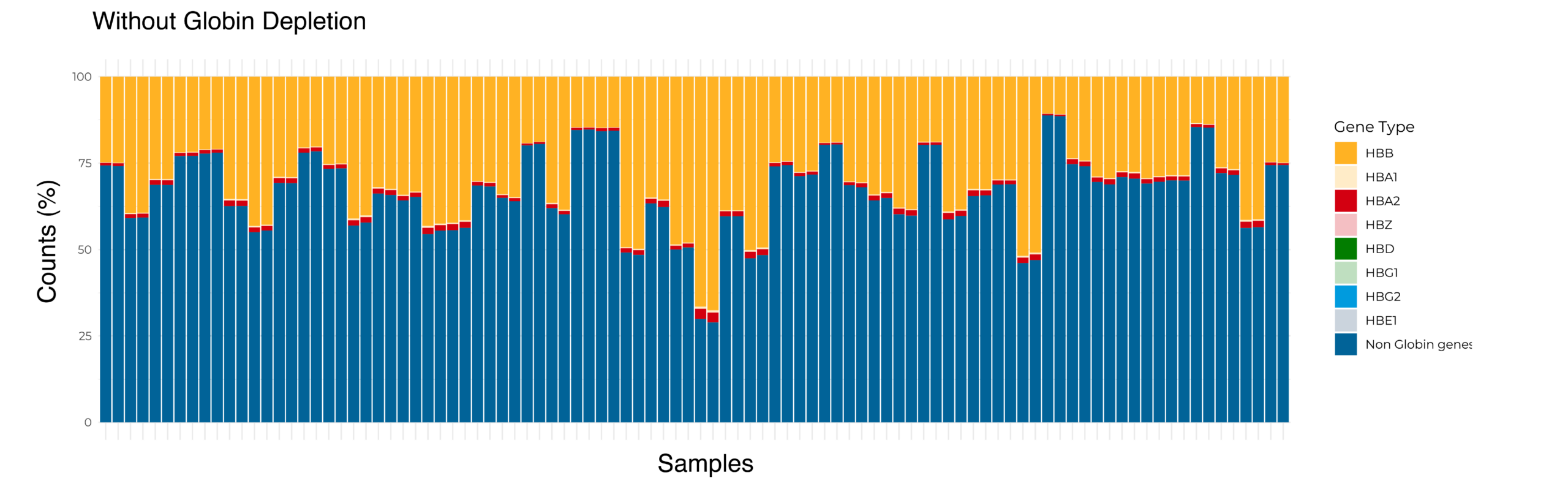
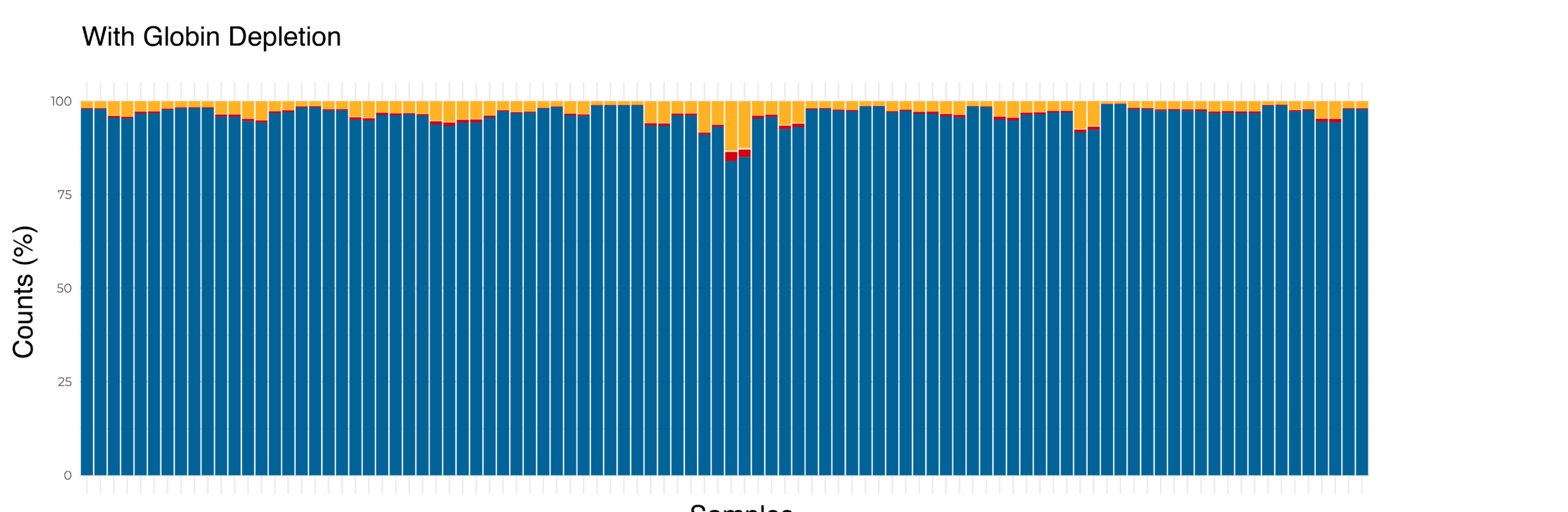
(Top) The distribution of reads mapped to non-globin and globin genes per sample for one 96 sample Total Blood BRB-seq library without globin depletion. An average of approximately one-third of reads mapped to globin genes. (Bottom) The distribution of reads mapped to non-globin and globin genes per sample for the same 96 samples in one Total Blood BRB-seq library treated with globin depletion. The reads associated with globin genes are drastically reduced, with more reads mapped to non-globin genes.
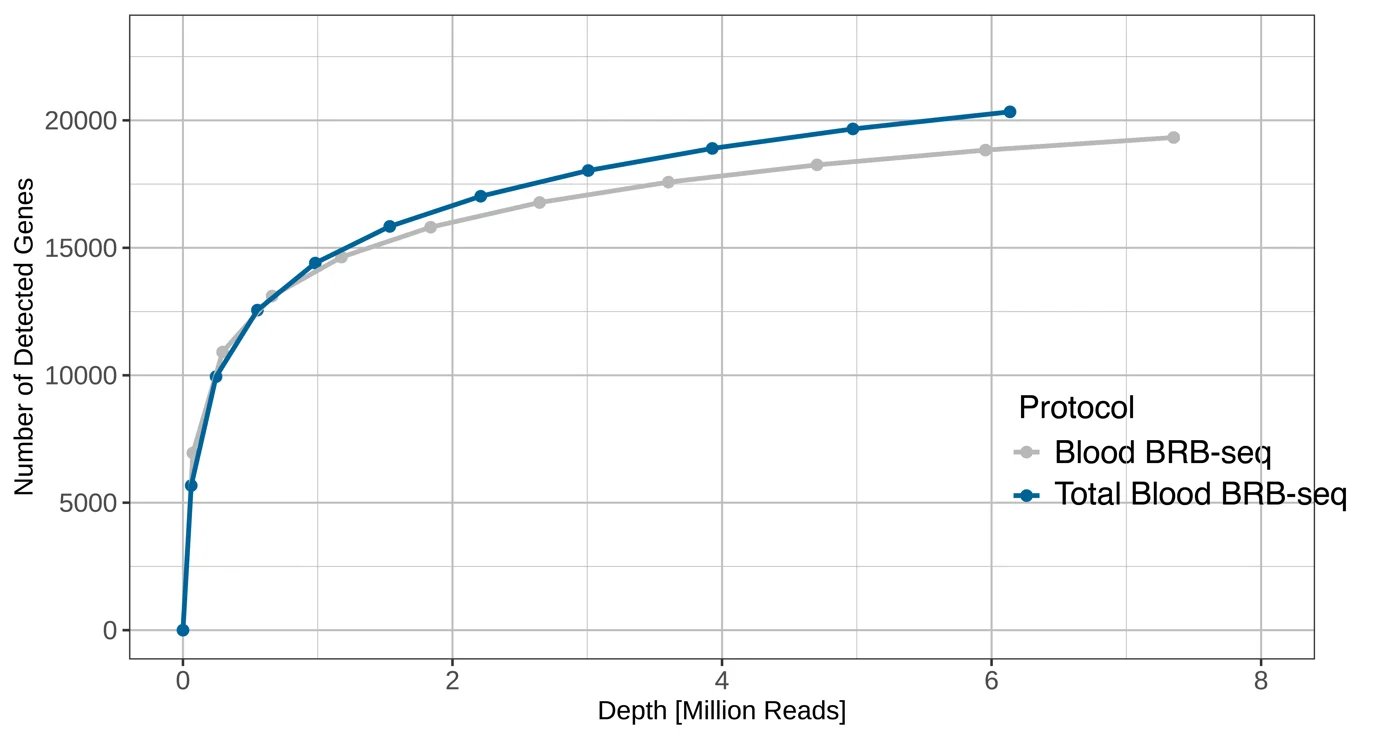
The plot shows the number of detected genes as a function of sequencing depth for both MERCURIUS™ Blood BRB-seq library preparation kit and the MERCURIUS™ Total Blood BRB-seq.
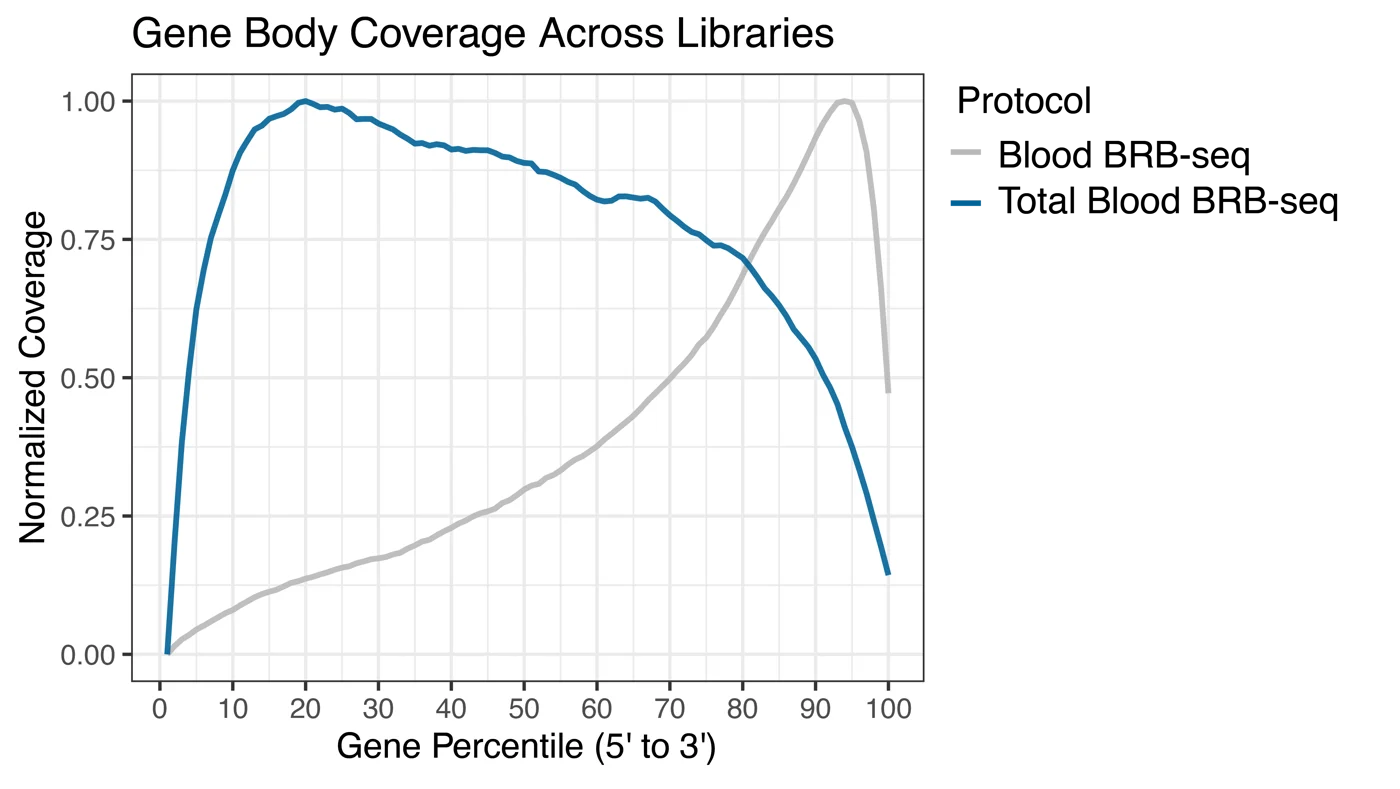
The gene body coverage shows a consistent and uniform read distribution across the entire gene body for the Total Blood BRB-seq protocol, while the Blood BRB-seq protocol shows a significant 3′ bias due to its poly-A selection methodology.
To have access to the deep-sequenced dataset (4.8 M reads per sample) contact us.
No, the Total Blood BRB-seq workflow is only dedicated to human blood samples.
For extracted RNA, we recommended a range of 10 ng-100 ng per sample.
The minimum recommended RIN number is 7 and the A260/280 ratio (Nanodrop) should be A260/280 >1.5.
We also offer high-throughput RNA extraction directly from PAXgene®, Tempus™ Blood RNA, and RNA shield tubes.
Total BRB-seq is a full-length total RNA sequencing method. We therefore normally recommend sequencing 2 to 10 million reads for each sample, which enables the reliable and unbiased detection of most of the genes.
As part of our standard service pipeline, we align the generated data to the genome of choice, provide a detailed report on the alignment and gene counting statistics and, finally, provide ready-to-use gene count matrices for downstream analysis.
You can either book a call with our experts to discuss your project or submit your experimental details via our contact form so we can review your design and requirements.
Based on the goals, sample types, and scale of your study, we may recommend starting with a pilot project to optimise conditions and de-risk a larger screen.
Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…