Discover Sample Datasets
PBMCs, HEK293
Built for teams requiring maximum scalability and sensitivity to detect the largest number of full-length genes possible from sorted single cells.
MERCURIUS™ Single-Cell FLASH-seq delivers ultra-sensitive full-length single-cell RNA sequencing (scRNA-seq) in an RNA-extraction-free 96- or 384-well plate-based workflow suitable for automation. Designed for maximum sensitivity, the technology detects ~2× more genes than other commercial solutions in a rapid one-day workflow. It enables sensitive, scalable scRNA-seq compatible with Illumina® and AVITI™ platforms.

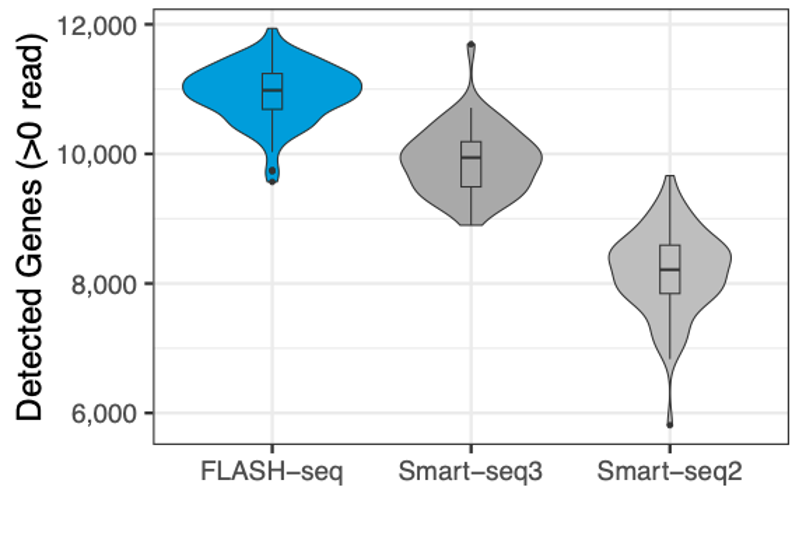
FLASH-seq detects a median of approximately 11,000 genes per sample in HEK293T cells, outperforming Smart-seq3 and Smart-seq2 after downsampling all to 500k reads/sample.
Gene body coverage shows a uniform read distribution across the entire gene body for the FLASH-seq protocol.
To have access to the deep-sequenced dataset (9.5M reads per sample) contact us.
We do not recommend very large cells, such as cardiomyocytes, or any other cell type incompatible with the FACS sorting step.
We have significantly enhanced the original FLASH-seq method to offer a streamlined workflow and superior data output. This plate-based technology (available in 96- and 384-well formats) features a novel, non-toxic tagmentation buffer and delivers ultra-sensitive gene detection, capturing up to two times more genes compared to other commercially available solutions.
We do not recommend very large cells, such as the cardiomyocytes, as they are not compatible with the FACS sorting step.
For optimal results, we require the cells to be directly FACS-sorted in the dedicated well plates. The 96- and 384-plates contain the lysis buffer. It is important for the user to sort the cells in the middle of the well. Please refer to the User Guide and the Sample Submission Guidelines for more details.
The average recommended sequencing depth is 250’000 reads/cell.
Product
Number of Samples
MERCURIUS™ Cell lysis module
MERCURIUS™ Cell lysis module
Single-cell RNA sequencing (scRNA-seq) has revolutionized our ability to examine the cellular heterogeneity of complex samples, identify new cell types, and explore transcriptional regulation at…
The Smart-seq2 protocol was the gold standard for full-length plate-based single-cell RNA-seq, offering superior sensitivity and the transcript coverage necessary to detect splice isoforms, allelic…
Lausanne, Switzerland and Frederick, MD – June 19, 2025 – Alithea Genomics, a pioneering biotech company based in Switzerland and the USA specializing in high-throughput…