Discover Sample Datasets
DLD1, SW837

Built for teams performing screening-scale high-throughput bulk RNA-seq on 3D spheroids and organoids seeking robust and reliable whole transcriptome insights for compound screens, safety assessment, dose-response, or mechanism of action studies.
MERCURIUS™ Spheroid DRUG-seq combines an optimized in-well spheroid lysis buffer with early sample barcoding and multiplexing for sensitive 3’ transcript coverage of coding transcripts in up to 384 samples in a single tube. The scalable, automation-ready one-day workflow is compatible with Illumina® and AVITI™ platforms without compromising depth, data quality, or sensitivity compared to sample-by-sample methods.
3D spheroid RNA-seq for high-throughput drug screening and dose-response studies in tumor- and organ-like models.
3D toxicology and safety assessment using transcriptome-wide readouts from more in vivo-like systems.
Comparing 2D vs 3D transcriptomic responses to prioritize compounds with better translational potential.
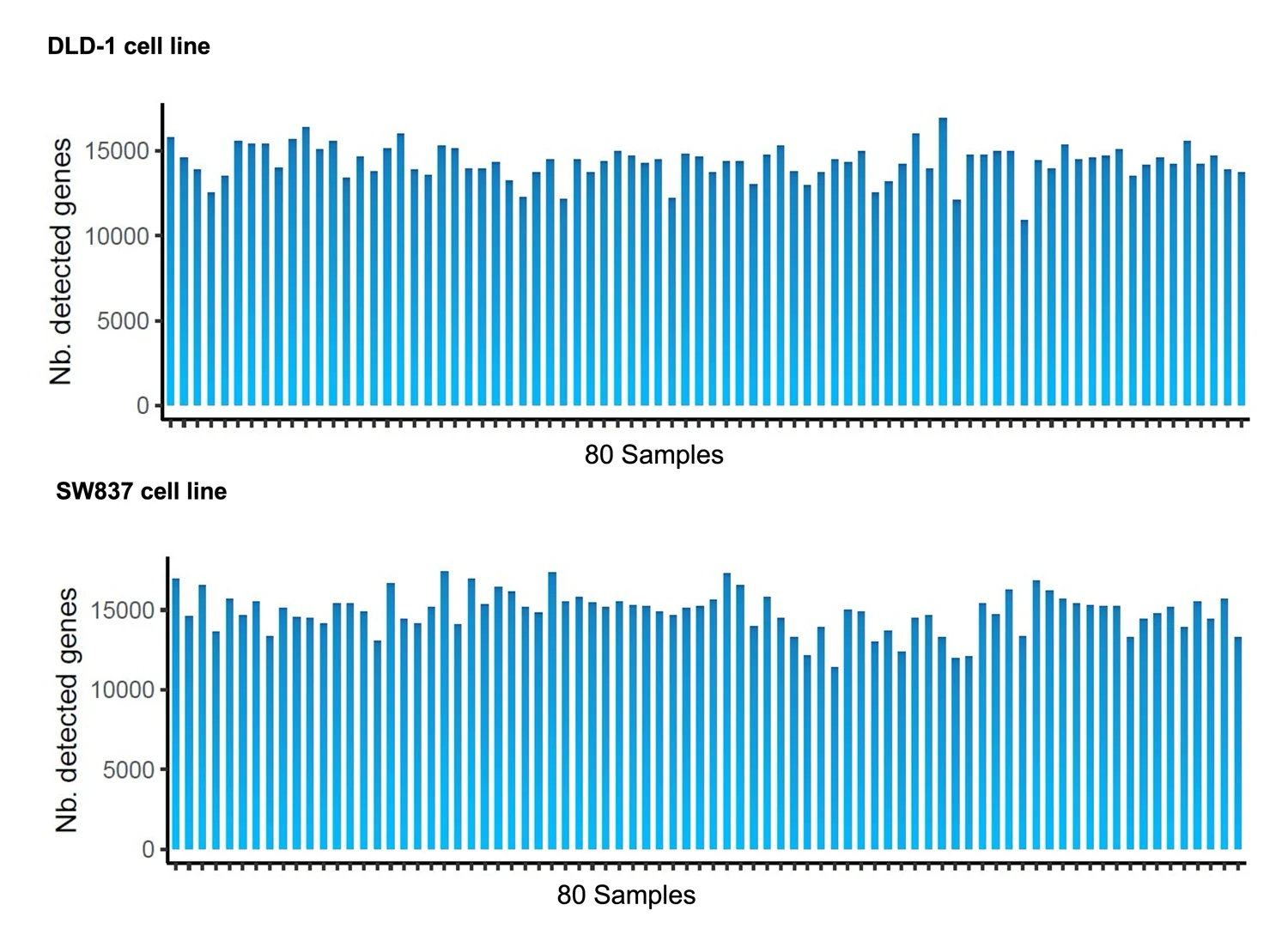
Number of genes detected per sample across 80-sample MERCURIUS™ Spheroid DRUG-seq runs in DLD-1 colorectal cancer spheroids (top) and SW837 colorectal cancer spheroids (bottom). An average of 13,000 genes is detected across all 80 samples from both cell lines at a sequencing depth of 1.7M reads/sample. The uniformity of detection across both cell lines and all samples demonstrates the reproducibility of MERCURIUS™ Spheroid DRUG-seq across different biological contexts.
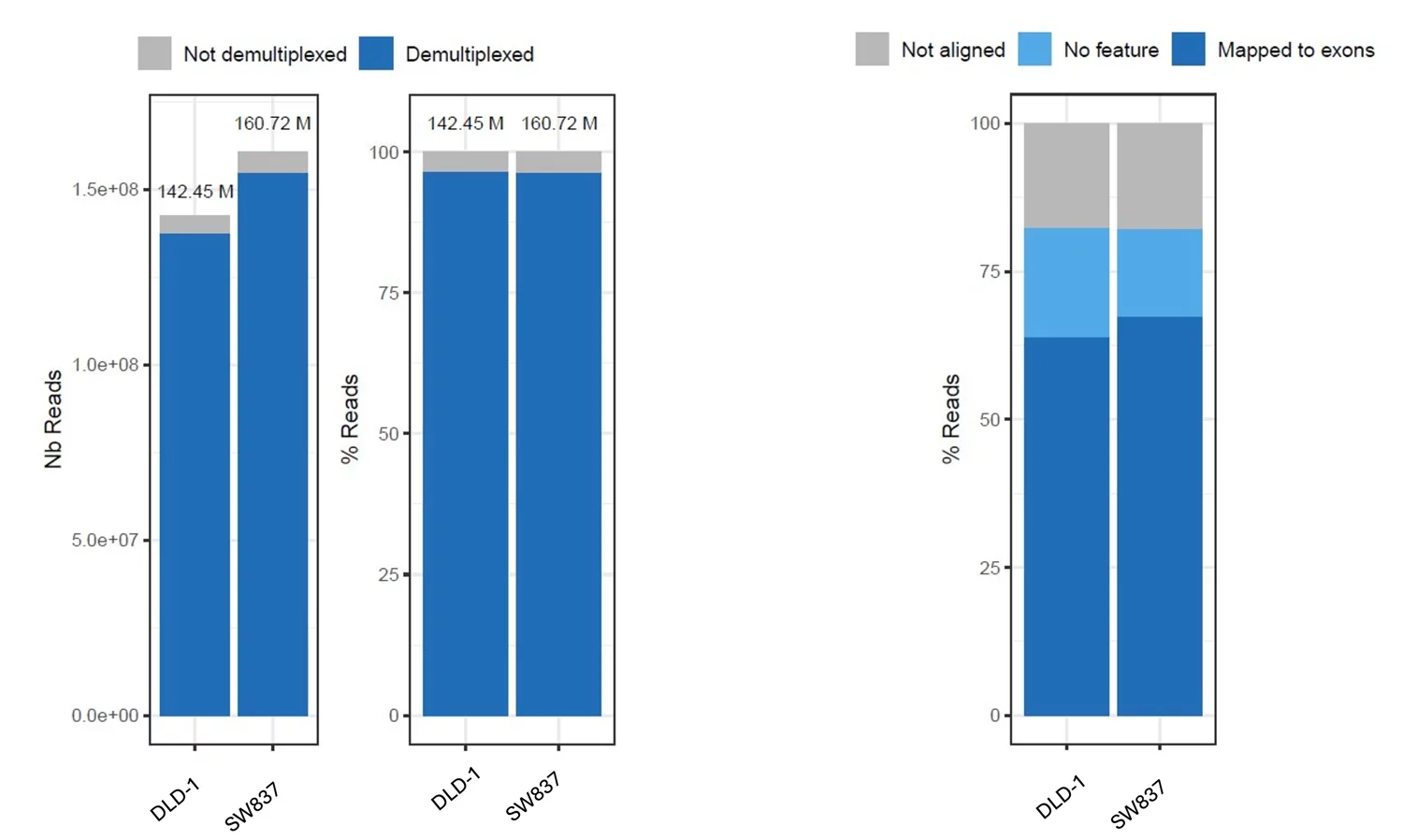
Sequencing quality metrics for MERCURIUS™ Spheroid DRUG-seq libraries prepared from spheroids of DLD-1 and SW837 cell lines. (Left) Demultiplexing efficiency expressed as the number of reads and the percentage of total reads, with both libraries achieving near-complete barcode assignment and minimal read loss. (Right) Mapping breakdown as a percentage of demultiplexed reads. Both libraries show comparable mapping profiles, with 80% of reads aligned, demonstrating consistent library quality across spheroid cell lines.
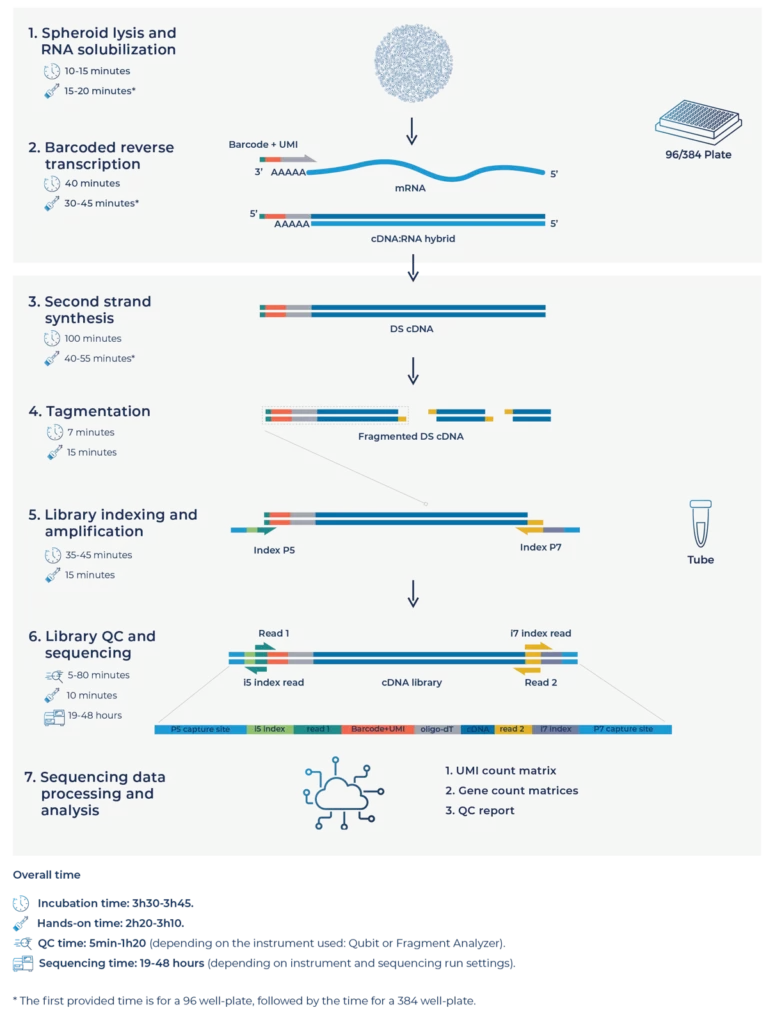
To have access to the deep-sequenced dataset (15G reads per sample) contact us.
To have access to the deep-sequenced dataset (17.5G reads per sample) contact us.
| Name | Origin | Species |
| U2OS | Osteosarcoma | Human |
| SWR837 | Adenocarcinoma | Human |
| DLD-1 | Adenocarcinoma | Human |
| 3D InSight™ Human Liver TOX Model | Liver | Human |
| 3D InSight™ Human Tumor Model | Liver | Human |
Each Spheroid DRUG-seq kit contains reagents (including four pairs of Unique Dual Indexing adapters) sufficient for the complete library preparation process for four different pools.
To note, the total number of RNA samples that can be processed with one kit does not exceed the kit specifications; for instance, a 96-sample kit can be used to prepare up to 96 samples distributed across up to four different libraries.
The recommended range of input material is of 2’000 – 50’000 cells per well.
The only difference between DRUG-seq and standard RNA-seq data analysis is the demultiplexing step, which is used to assign sequencing reads to their sample of origin based on the Spherioid DRUG-seq barcode sequence.
For a thorough description of Spheroid DRUG-seq data processing, please refer to the kit user guide.
The barcode set for your kit is conveniently located on the kit label. Please refer to the label for accurate identification.
For optimal compatibility, ensure that you use the appropriate plate format (e.g., for kits designed for 96 reactions, the 96 well-plate format should be used). This ensures accurate and efficient processing of your samples. If you have any further questions or concerns, please contact our support team for assistance by email or using our live chat tool.
Product
Number of Samples
MERCURIUS™ Spheroid Cell Lysis Module
MERCURIUS™ UDI Expansion Module
MERCURIUS™ Standard Post-Pooling Preparation Module (4 libraries)
Each kit includes 4 UDI pairs. Add the expansion module if you need more unique indexes (total 16 UDI pairs available).
Biomarker discovery using gene expression signatures has transformed pharmaceutical discovery and development, from early hit triage and target validation to toxicology, dose optimization, and patient…
Next-generation AI models for drug discovery are only as powerful as the quality, size, and perturbation-richness of their training data. Yet no publicly available dataset…
As the SOT Annual Meeting and ToxExpo 2026 approaches, we take a pre-conference look at the program through a transcriptomics-focused lens. One overall shift is…
Identifying brain-penetrant small-molecule modulators of human microglia using a cellular model of synaptic pruning – McCrea and Batorsky et al. [1] Published May 9th, 2025,…