Built for teams aiming to detect coding and non-coding transcripts, isoforms, and splice variants in purified RNA from larger cohorts, more conditions, or more timepoints than possible with standard total RNA-seq methods.
MERCURIUS™ Total BRB-seq combines early sample barcoding and multiplexing for full-length transcript coverage of coding and non-coding transcripts in 96 purified RNA samples in a single tube. It enables high-throughput total RNA-seq on Illumina® and AVITI™ platforms without compromising depth, data quality, or sensitivity compared to sample-by-sample methods.
Total RNA bulk RNA-seq when both mRNA and non-coding RNA signatures are required (e.g. toxicogenomics, regulatory RNA).
Mechanistic and pathway analysis where long non-coding RNAs, antisense transcripts and other RNA classes contribute to phenotype.
Building richer transcriptomic datasets for biomarker discovery and ML/AI models from purified RNA.

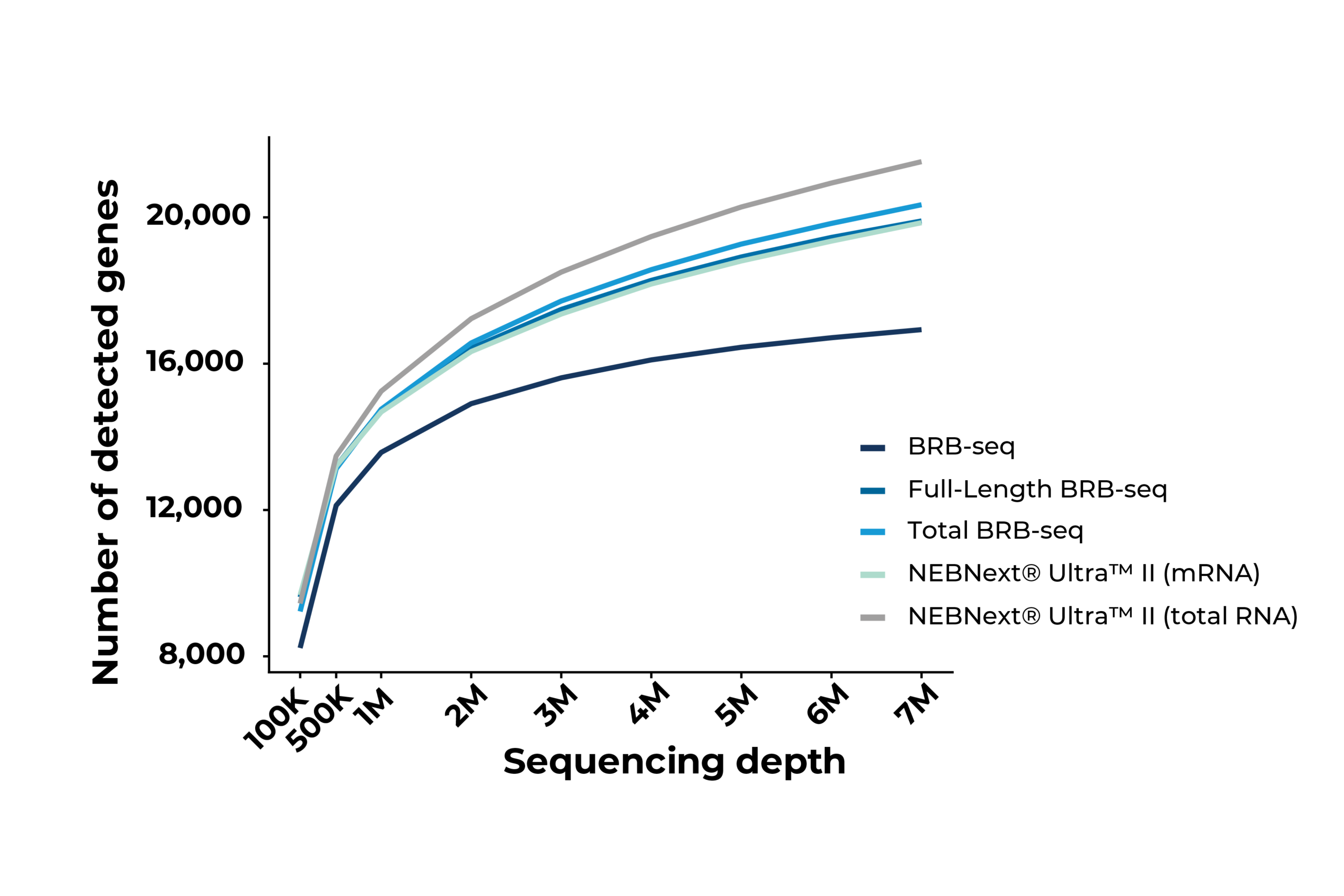
Saturation curves indicating the number of detected genes as a function of sequencing depth in Huh7 cells. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq perform comparably to NEBNext® Ultra™ II mRNA and Total RNA preparations, respectively, demonstrating equivalent sensitivity at a fraction of the library preparation cost and effort. MERCURIUS™ BRB-seq is a 3’ mRNA library preparation technology, hence the lower overall gene detection versus full-length methods.

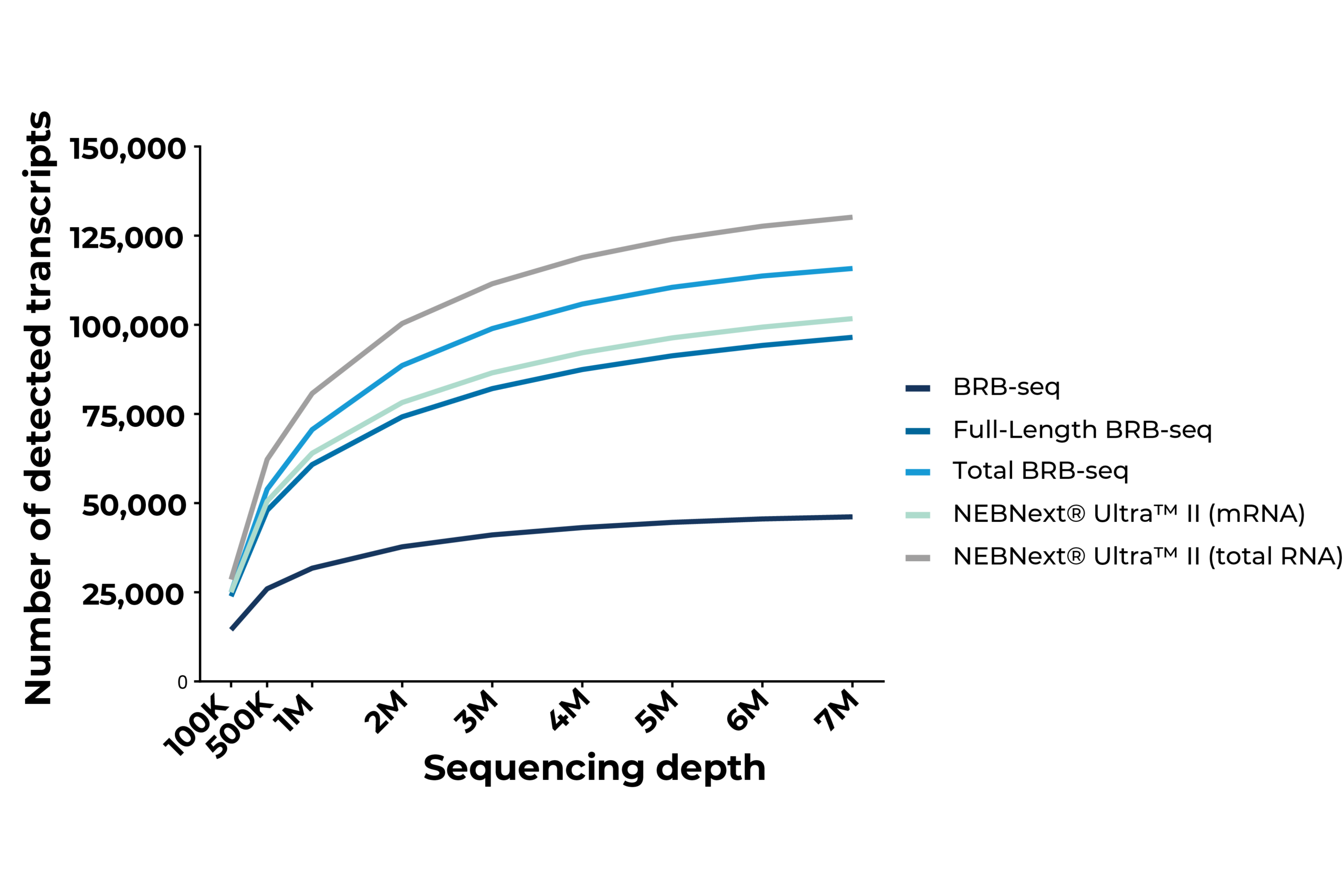
Saturation curves indicating the number of detected transcripts as a function of sequencing depth in Huh7 cells. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq perform comparably to NEBNext® Ultra™ II mRNA and Total RNA preparations, respectively, demonstrating equivalent sensitivity at a fraction of the library preparation cost and effort. MERCURIUS™ BRB-seq specifically detects protein-coding transcripts, hence the lower overall transcript detection compared to full-length methods.
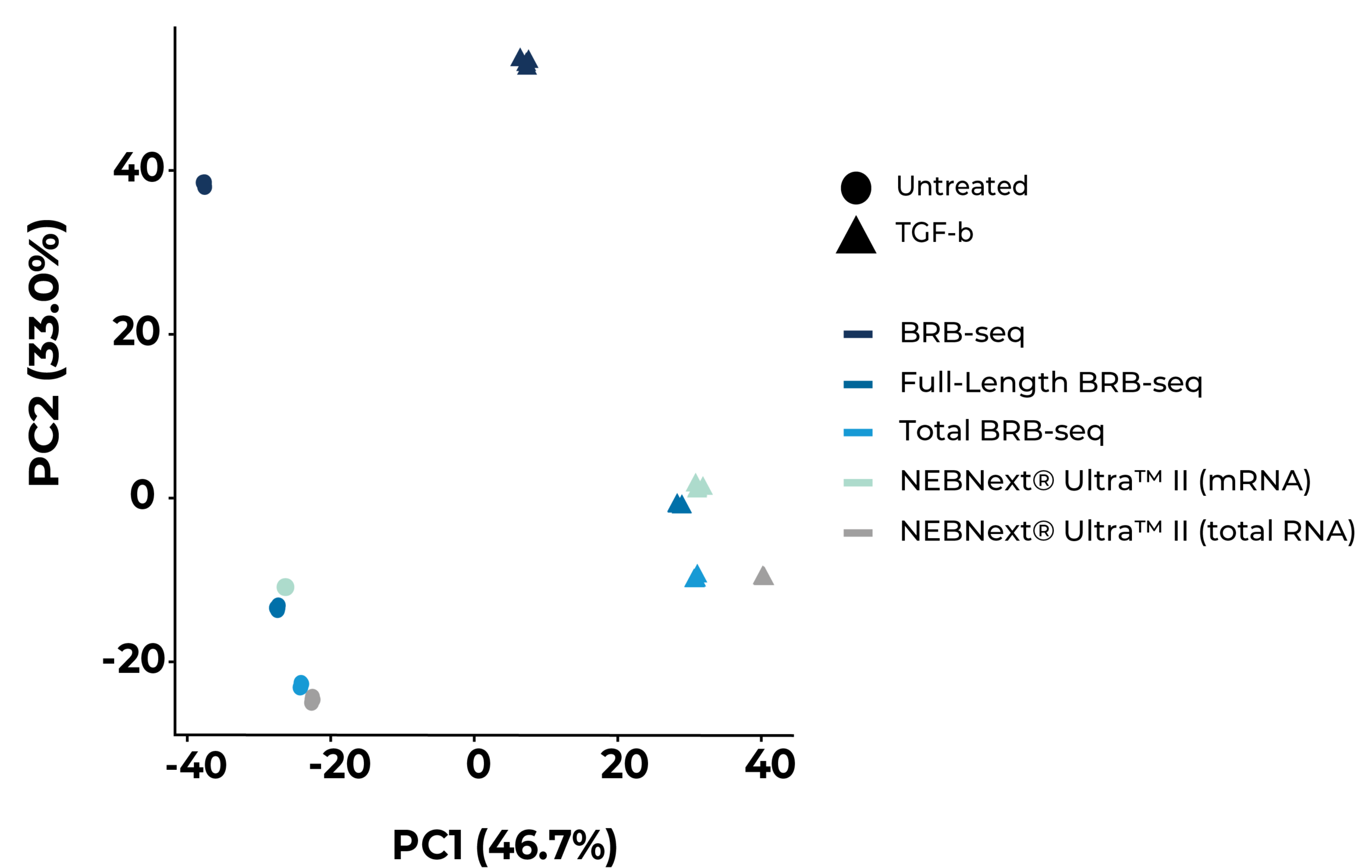
Principal component analysis (PCA) of gene expression profiles generated by the specified library preparation methods across untreated (circles) and TGF-β-treated (triangles) Huh7 samples. Samples cluster primarily by library type along PC2 and by treatment condition along PC1. All methods robustly capture the expression response to treatment. MERCURIUS™ Full-Length BRB-seq and Total BRB-seq cluster closely with their NEBNext® Ultra™ II counterparts, demonstrating equivalent transcriptomic profiles. MERCURIUS™ BRB-seq clusters separately, consistent with its 3′-end capture design.
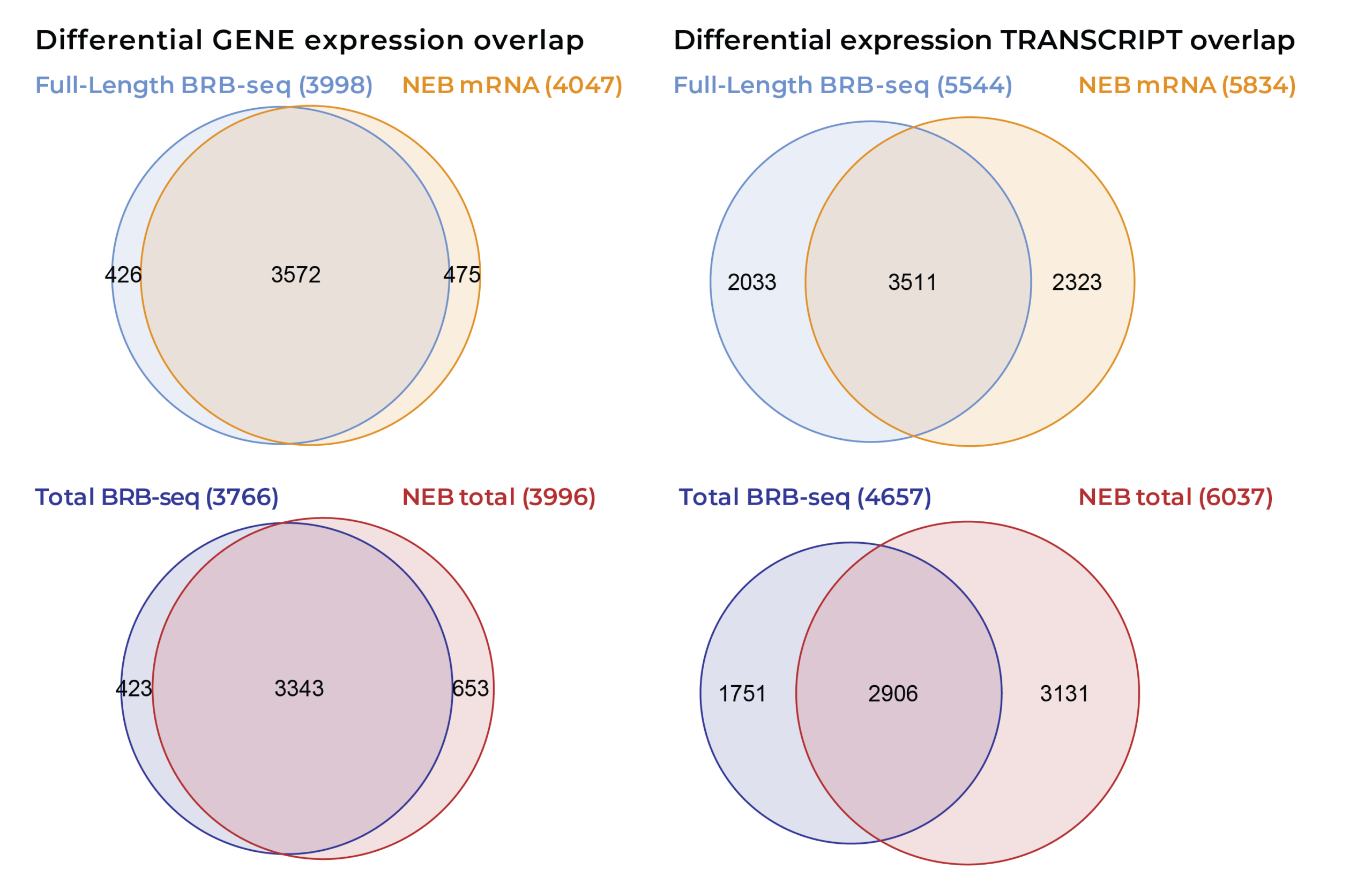
Venn diagrams showing the overlap in differentially expressed (DE) genes (left) and transcripts (right) identified by MERCURIUS™ Full Length BRB-seq and NEBNext® Ultra™ II for mRNA libraries (top) and total RNA libraries (bottom). Numbers indicate DE features unique to each method or shared between both. At the gene level, MERCURIUS™ Full Length BRB-seq and NEBNext® Ultra™ II share over 88% of DE genes for mRNA and 84% for total RNA preparations, demonstrating strong concordance in differential expression calls across both library types. The overlap is smaller but still strong for DE transcripts across technologies.
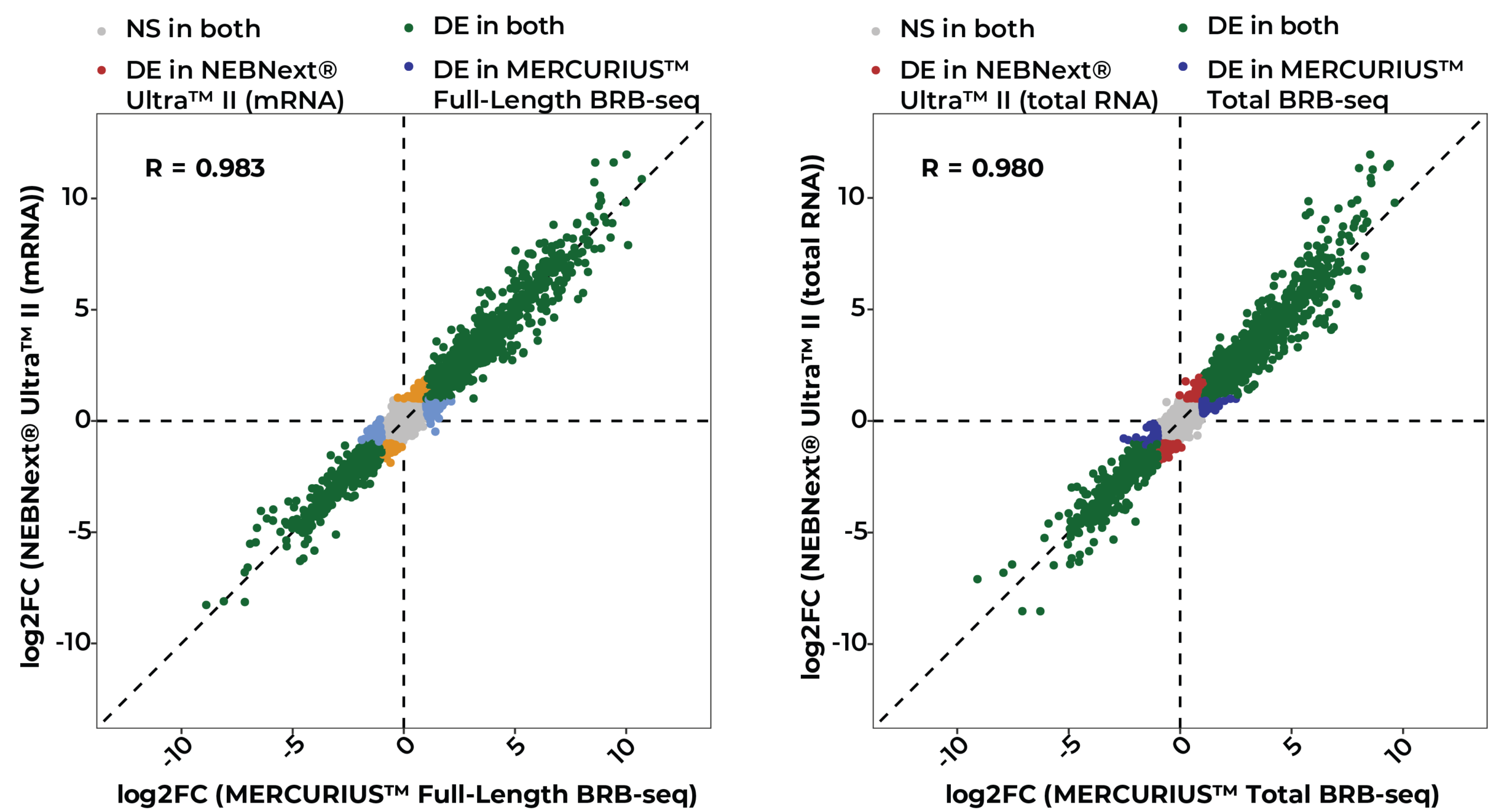
Log2 fold-change correlation between MERCURIUS™ Full-Length BRB-seq and NEBNext® Ultra™ II for mRNA (left, R = 0.983) and total RNA (right, R = 0.980) protocols, across intersecting gene sets of 13,364 and 13,237 genes, respectively. Each point represents a gene, colored by its differential expression (DE) or non-significant (NS) status after treatment of Huh7 cells with human TGF-β versus untreated cells. The dashed diagonal represents perfect correlation. The concordance of DE fold changes across both RNA input types demonstrates that MERCURIUS™ Full-Length and Total BRB-seq reliably recapitulate the results of gold-standard library preparation.
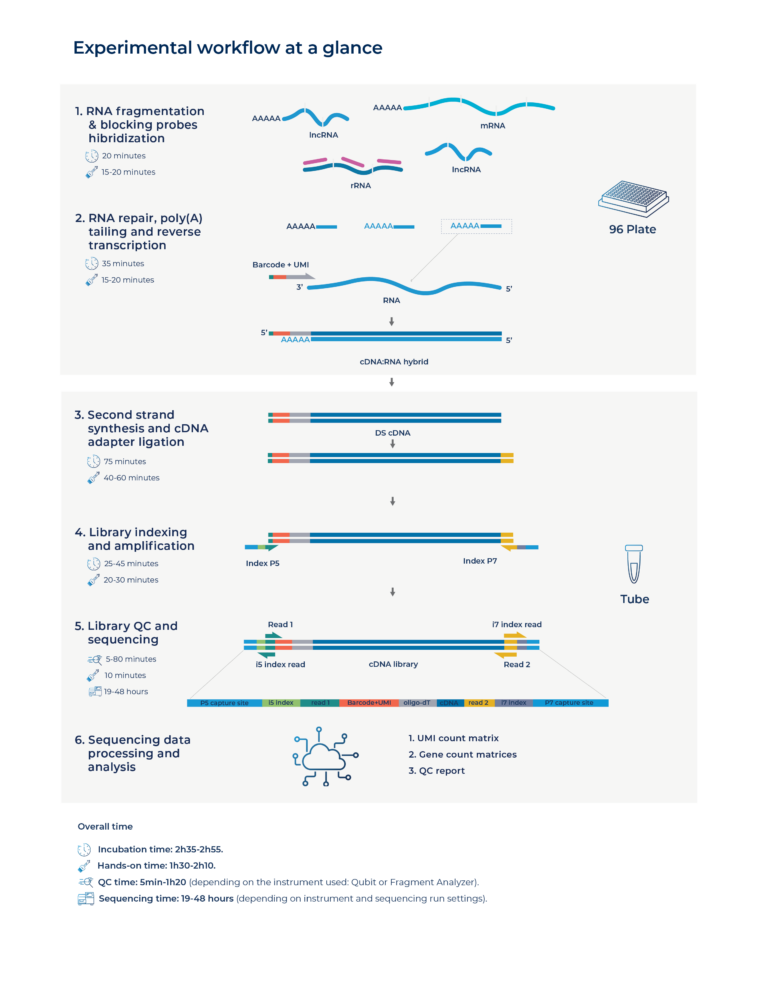
Each kit includes 4 Unique Dual Index (UDI) pairs, so you can prepare up to 4 indexed libraries (pools) per kit.
The tested input range is 100 pg to 100 ng total RNA per well. For best results (library complexity + uniformity), aim for 10–100 ng per well, and pool at least 8 wells per library.
Because Total BRB-seq uses early multiplexing, it’s important that RNA quantity and quality are consistent across samples (e.g., RIN > 7, 260/230 > 1.5, and ~±10% input uniformity).
Total BRB-seq is designed for total RNA sequencing (not just mRNA), including coding and non-coding transcripts, and includes an rRNA depletion step prior to library prep.
We recommend 10-20 million reads for each sample, which enables the detection of 65,000 transcripts.
Yes, there is one extra step: after standard index demultiplexing, you demultiplex by sample barcodes to generate per-sample gene/transcript count outputs. The workflow supports producing outputs like UMI count matrices, gene count matrices, and a QC report.
Go to the “Resources” section of this page and click on the button to download the barcode files for the product you are using.
Yes. If you are not ready to run kits internally, Alithea can run the workflow and deliver FASTQ files, gene count matrices, and analysis report files.
* Contact us to inquire about compatibility with other species.
Product
Catalog Number
MERCURIUS™ UDI X-Leap Expansion module
MERCURIUS™ Full-Length Post-Pooling Preparation Module (4 libraries)
Each kit includes 4 UDI pairs. Add the expansion module if you need more unique indexes (total 16 UDI pairs available).
Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…