Discover Sample Datasets
PBMCs, HEK293
Built for teams requiring maximum sensitivity to detect the largest number of full-length genes possible from precious, ultra-low-input RNA samples (1 pg – 1 ng per well).
The MERCURIUS™ Low-Input FLASH-seq RNA-extraction-free protocol combines ultra-sensitive gene detection with full-length transcript coverage in a 96-well plate-based workflow. It enables ultra-low-input RNA-seq on Illumina® and AVITI™ platforms without compromising depth, data quality, or sensitivity.
Low-input RNA-seq for scarce samples (e.g. small biopsies, rare or sorted populations, limited clinical material) where standard RNA-seq kits fail.
Full-length mRNA sequencing of low-input RNA to profile differential gene expression, splicing variants and isoforms from precious samples at pg to ng input.

FLASH-seq detects a median of approximately 11,000 genes per sample in HEK293T cells, outperforming Smart-seq3 and Smart-seq2 after downsampling all to 500k reads/sample.
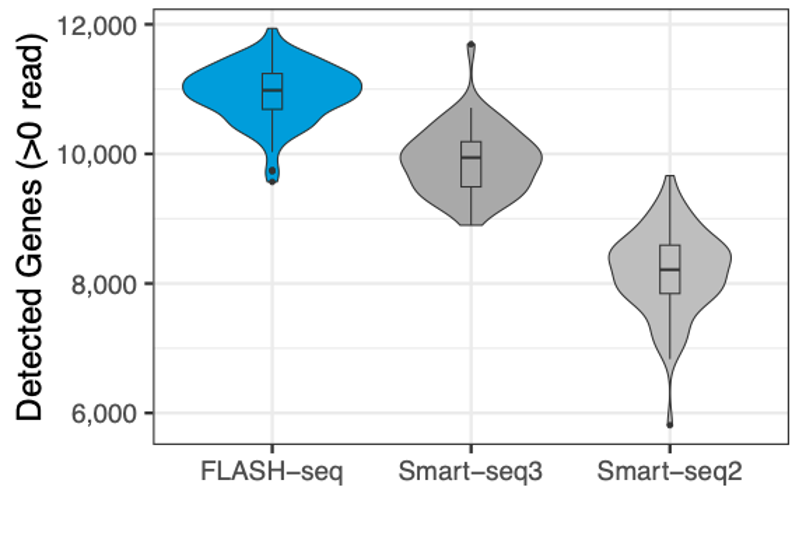
Number of genes detected in human PBMCs, processed with different protocols, and the number of reads downsampled to 125,000 raw reads.
Gene body coverage shows a uniform read distribution across the entire gene body for the FLASH-seq protocol.
The number of genes detected in HEK 293T cells using different RNA inputs (from 2.5 pg to 250 pg) at different sequencing depths.
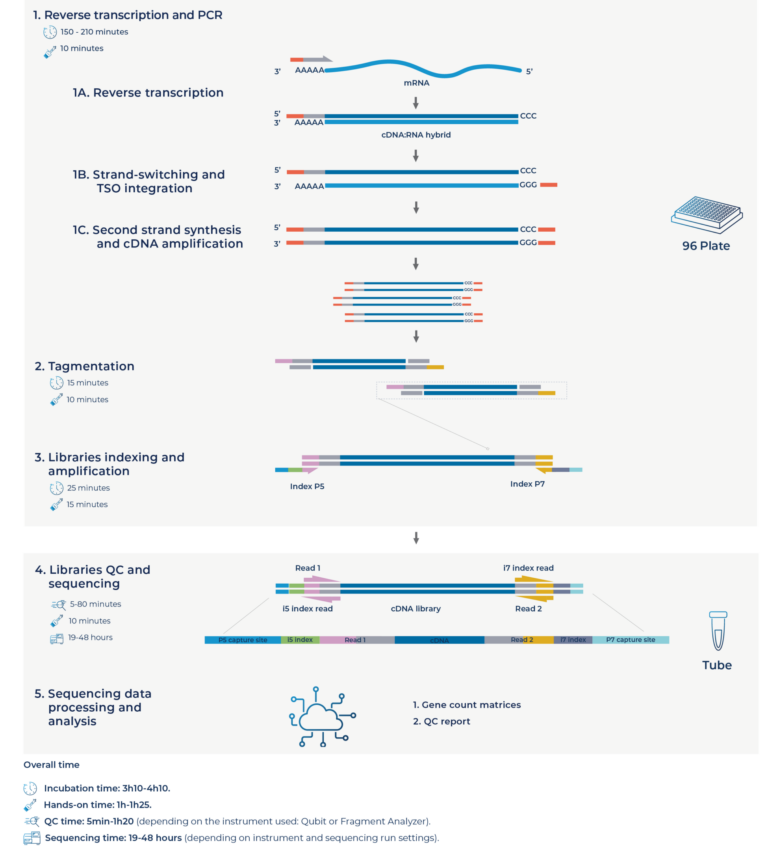
To have access to the deep-sequenced dataset (9.5M reads per sample) contact us.
We have significantly enhanced the original FLASH-seq method to offer a streamlined workflow and superior data output. This 96-well plate-based technology features a novel, non-toxic tagmentation buffer and delivers ultra-sensitive gene detection, capturing up to twice as many genes as other commercially available solutions.
The MERCURIUS™ Low-Input FLASH-seq protocol is designed for low RNA input from precious samples (1 pg to 1 ng), subcellular inputs, liquid biopsies, exosomes, microbiopsies, small organoids, and early embryos.
You are required to extract RNA from the samples and normalize them to the same concentration prior to freezing and shipping.
Please note! To preserve sample quality during shipping, plates must be placed between dry ice—not just on top. Inadequate cooling may compromise RNA integrity!
The average recommended sequencing depth is 250’000 reads/cell.
Single-cell RNA sequencing (scRNA-seq) has revolutionized our ability to examine the cellular heterogeneity of complex samples, identify new cell types, and explore transcriptional regulation at…
The Smart-seq2 protocol was the gold standard for full-length plate-based single-cell RNA-seq, offering superior sensitivity and the transcript coverage necessary to detect splice isoforms, allelic…
Lausanne, Switzerland and Frederick, MD – June 19, 2025 – Alithea Genomics, a pioneering biotech company based in Switzerland and the USA specializing in high-throughput…
Revolutionizing Single-Cell RNA Sequencing with FLASH-seq The field of genomics is advancing rapidly, and one of the most transformative tools has been single-cell RNA sequencing…