This service enables full-length mRNA transcript coverage from low RNA inputs from precious samples (1pg to 1ng).
The MERCURIUS™ Low-input FLASH-seq service offers the most sensitive, full-length mRNA and cost-effective solution even for low-abundance genes, empowering researchers to explore differential gene expression, detect alternative splicing, and analyze isoform diversity, all critical for understanding complex biology, especially in rare cell populations.
This service is dedicated to low RNA input from precious samples (1 pg to 1 ng), subcellular inputs, liquid biopsies, exosomes, microbiopsies, small organoids, and early embryos.
As part of the FLASH-seq service, clients simply need to ship the frozen plates with the samples to our service centers located in Switzerland and the United States.
As a result of our service, our clients receive raw sequencing data (fastq files), gene count matrices, and analysis report files.

2 days
(Qubit, Fragment analyzer, shallow sequencing)
1 week
1 week
Raw FASTQ files, sample report file, QC files, and gene count tables
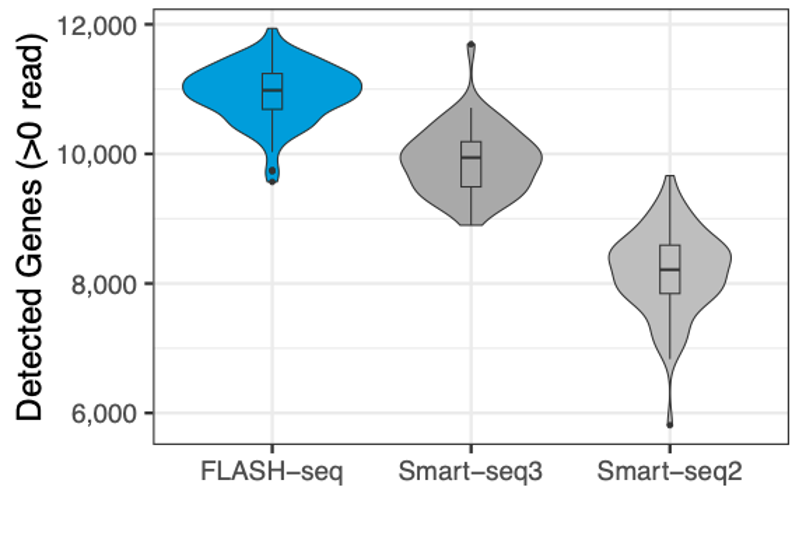
FLASH-seq detects a median of approximately 11,000 genes per sample in HEK293T cells, outperforming Smart-seq3 and Smart-seq2 after downsampling all to 500k reads/sample.
Number of genes detected in human PBMCs, processed with different protocols, and the number of reads downsampled to 125,000 raw reads.
Gene body coverage shows a uniform read distribution across the entire gene body for the FLASH-seq protocol.
The number of genes detected in HEK 293T cells using different RNA inputs (from 2.5 pg to 250 pg) at different sequencing depths.
To have access to the deep-sequenced dataset (7.3 M reads per sample) contact us.
To have access to the deep-sequenced dataset (7.1 M reads per sample) contact us.
We have significantly enhanced the original FLASH-seq method to offer a streamlined workflow and superior data output. This plate-based technology (available in 96- and 384-well formats) features a novel, non-toxic tagmentation buffer and delivers ultra-sensitive gene detection, capturing up to two times more genes compared to other commercially available solutions.
The MERCURIUS™ Low-Input FLASH-seq protocol is designed for low RNA input from precious samples (1 pg to 1 ng), subcellular inputs, liquid biopsies, exosomes, microbiopsies, small organoids, and early embryos.
You are required to extract RNA from the samples and normalize them to the same concentration prior to freezing and shipping.
Please note! To preserve sample quality during shipping, plates must be placed between dry ice—not just on top. Inadequate cooling may compromise RNA integrity!
From the moment we receive the frozen plates until the moment we deliver the sequencing data, we typically estimate four weeks.
You can either book a call with our experts to discuss your project or submit your experimental details via our contact form so we can review your design and requirements.
Based on the goals, sample types, and scale of your study, we may recommend starting with a pilot project to optimise conditions and de-risk a larger screen.
If you’re interested in implementing the technology in your own lab instead, you can explore our MERCURIUS™ Low-Input FLASH-seq kits on the dedicated kits page.
Single-cell RNA sequencing (scRNA-seq) has revolutionized our ability to examine the cellular heterogeneity of complex samples, identify new cell types, and explore transcriptional regulation at…
The Smart-seq2 protocol was the gold standard for full-length plate-based single-cell RNA-seq, offering superior sensitivity and the transcript coverage necessary to detect splice isoforms, allelic…
Revolutionizing Single-Cell RNA Sequencing with FLASH-seq The field of genomics is advancing rapidly, and one of the most transformative tools has been single-cell RNA sequencing…