Massively multiplexed and full-length RNA-seq directly on cell, organoid and tissue lysates - no need for prior RNA extraction.
Discover Sample Datasets
Human
96 samples in a single tube
From differential gene expression to splicing variants and isoform detection
Prepare libraries directly from frozen cell pellets.
The more samples processed, the lower the cost per sample.
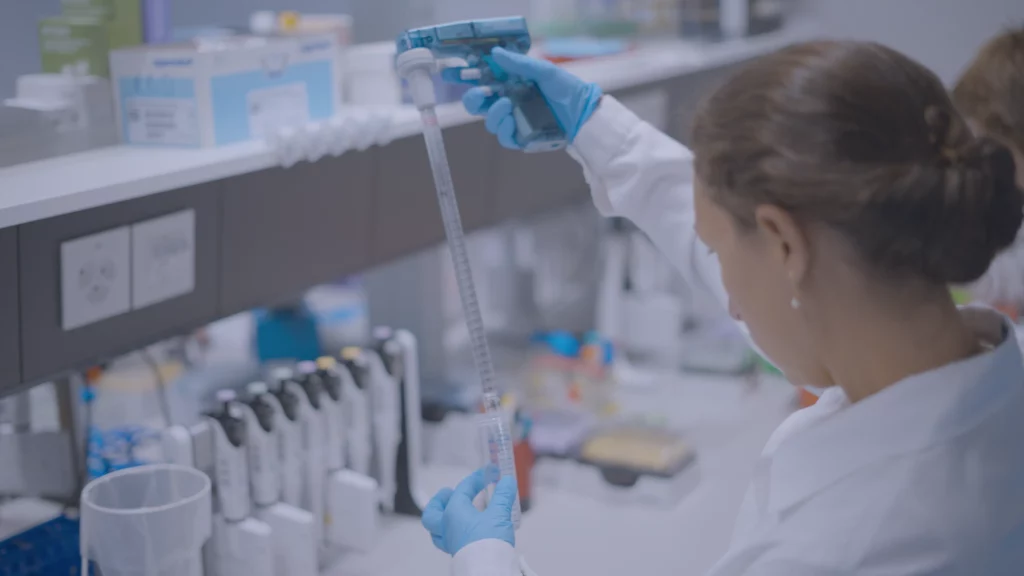
The MERCURIUS™ Full-Length DRUG-seq service is a convenient and scalable solution for projects of any size, enabling transcriptome-wide detection of full-length coding transcripts.
Users simply deliver their 96-well plates containing frozen cells to us in Switzerland or the United States.
Upon receipt of the plates, our team prepares the libraries, then sequences to your desired read depth per sample, and then performs data pre-processing.
We return results, including raw fastq files, sequencing and alignment reports, and gene count matrices suitable for downstream differential expression analyses, once the data meet our rigorous quality control criteria.
During the process, we always keep clients informed at defined checkpoints so we can decide together how best to proceed to the next steps.

2 days
(Qubit, Fragment analyzer, shallow sequencing)
1 week
1 week
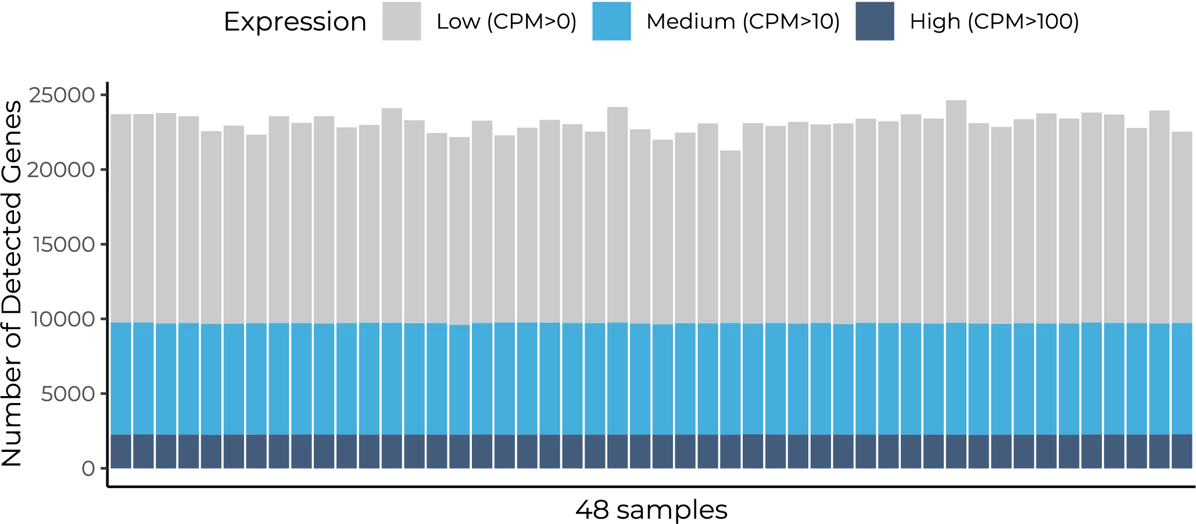
Distribution of the number of detected genes across 48 samples prepared with the MERCURIUS™ Full-Length BRB-seq service. The library was sequenced at an average of 12 million reads.
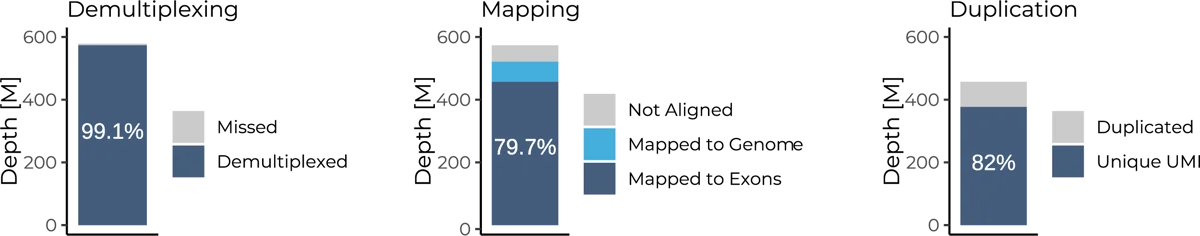
Full-Length BRB-seq performance shows 99% demultiplexing rate from raw data, 78% mapping rate, and 18% duplication rate of the 48 pooled samples and sequenced at 12 million reads per sample.
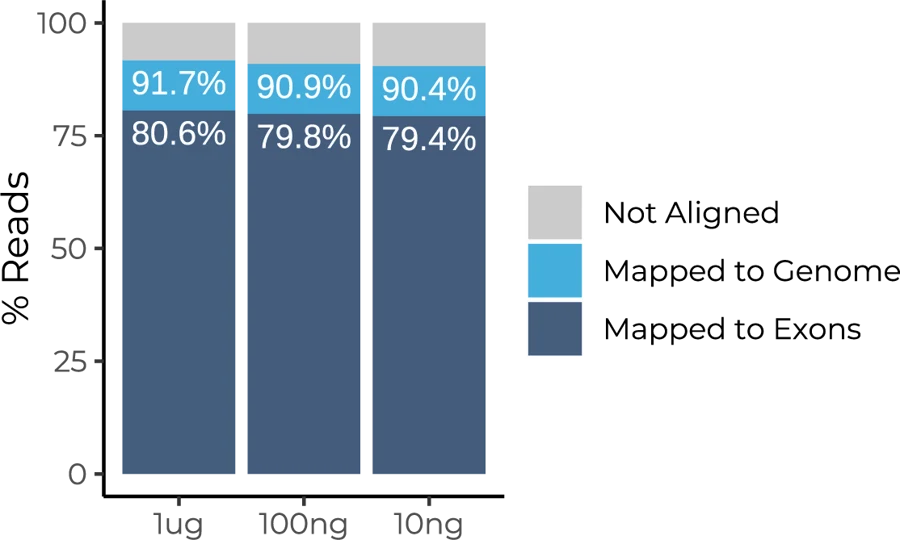
Mapping rates are uniform across three RNA inputs tested: 1ug, 10ng, and 100ng. Human Universal Reference RNA was used to prepare mRNA Full-Length BRB libraries.
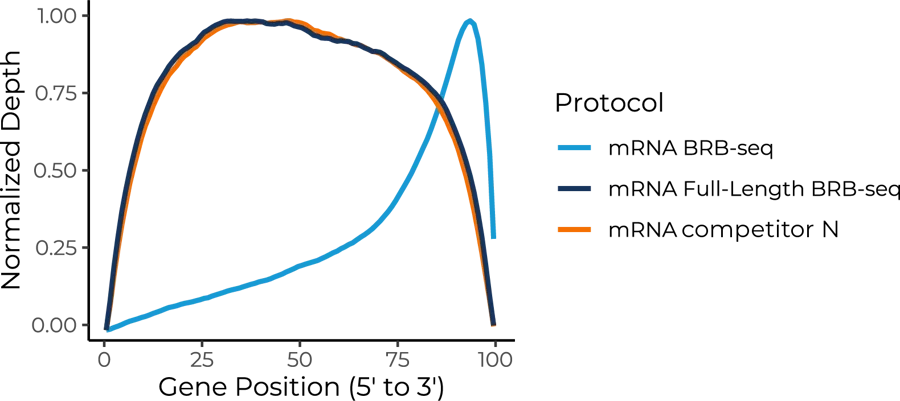
The gene body coverage shows a consistent and uniform reads distribution across the entire gene body for the Full-Length BRB-seq protocol, comparable to competitor N protocol, while the BRB-seq protocol shows a significant 3′ bias due to its poly-A selection methodology.
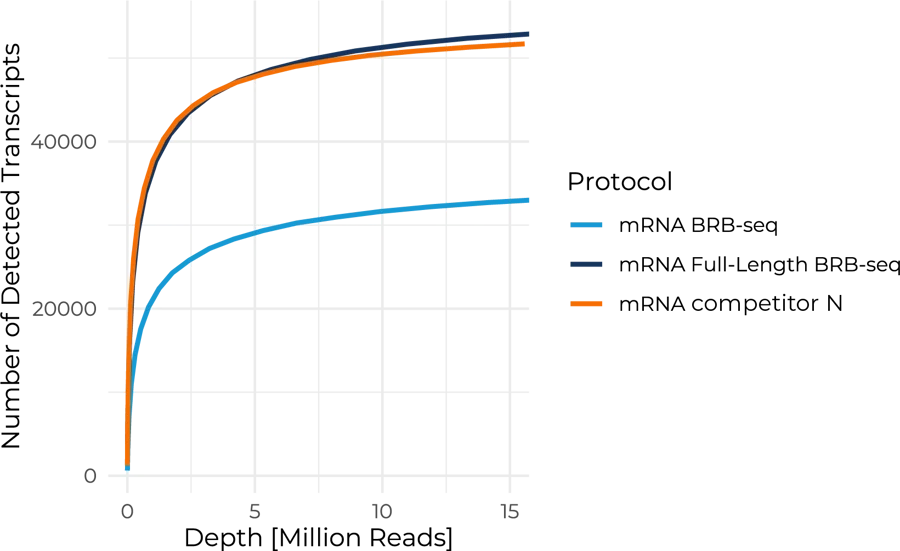
Benchmarking the Full-Length BRB-seq protocol against competitor N shows similar performance for isoform detection. More than 50,000 isoforms can be detected at a sequencing depth of 12M reads per sample.
To have access to the deep-sequenced dataset contact us.
To generate high-quality sequencing data we recommend starting with 5’000 – 25’000 mammalian cells per well for 96 well-plate and 2’000 – 10’000 cells per well for 384 well-plate.
The total RNA amount per pool should be at least 80’000 cells.
MERCURIUS™ DRUG-seq can be performed on various sample types, including: Cell lines, Primary cells, Organoids, or Spheroids.
Check out our DRUG-seq Kits page for a list of our validated cell lines.
Full-Length DRUG-seq provides comprehensive coverage of the full-length mRNA transcripts.
We, therefore, normally recommend sequencing 5-20 million reads for each sample, depending on project scale and scope.
You can either book a call with our experts to discuss your project or submit your experimental details via our contact form so we can review your design and requirements. Based on the goals, sample types, and scale of your study, we may recommend starting with a pilot project to optimise conditions and de-risk a larger screen.
Determining the most suitable transcriptomic technology to drive your large-scale compound screen, clinical study, or to assess a panel of genetic perturbations can be a…
High-throughput’ in sequencing refers to the amount of DNA molecules read at the same time. Technologies are now capable of sequencing many fragments of DNA…
With a growing number of published 3’ mRNA-seq methods now available, researchers have more choices than ever for high-throughput and cost-effective transcriptomic screening. While broadly…