Next-generation sequencing (NGS) has now entered an era of unprecedented scalability. Two companies, Illumina and MGI Tech, lead the way in the ultra-high-throughput chapter of NGS. Illumina and MGI sequencing technologies produce a significant amount of data and allow researchers to simultaneously process more samples at a lower cost than ever before.
In this article, we tease apart these technologies and discuss the scalability of the most recent genetic sequencers from Illumina and MGI.
What is scalability?
Scalability refers to the ability to process large numbers of samples in a single sequencing run.
To do so, samples are barcoded with unique indices to differentiate between samples. This is known as multiplexing.
The number of samples you can multiplex depends on two main factors: the number of unique indices available, and the desired sequencing depth per sample.
As sequencing platforms produce more data, more samples can be processed simultaneously.
Comparison of Illumina and MGI sequencing technologies
Illumina and MGI sequencing technologies are short-read sequencing by synthesis approaches with high scalability and comparable sequencing results, however, the methods used have some significant differences (Mortazavi et al., 2008; Drmanac et al., 2010; Goodwin, McPherson and McCombie, 2016).
Bridge amplification versus rolling circle amplification and DNA nanoballs
Illumina sequencing uses bridge amplification to amplify DNA fragments on a solid support. In contrast, MGI sequencing uses rolling circle amplification (RCA) in solution to generate long tightly coiled single strands of head-to-tail DNA known as DNA nanoballs (DNBs) (Drmanac et al., 2010).
The main advantage of RCA is that the original copy of DNA acts as the template for each new copy. This process reduces amplification errors common in PCR-based bridge amplification (Drmanac et al., 2010).
Furthermore, MGI sequencing uses patterned flow cells containing positively charged spots placed at optimal distances for maximum data generation. The binding of DNBs to the flow cell is highly efficient thanks to their negative charge combined with their defined size.
Sequencing by synthesis
The sequencing by synthesis methods used by Illumina and MGI sequencing are similar but with some key differences.
In Illumina and some older MGI options, fluorescently labeled nucleotides are added, and the emission wavelength from each cluster is recorded and decoded to identify the base. These are then cleaved to incorporate the next base (Goodwin, McPherson and McCombie, 2016).
MGI has recently developed an innovative variation of sequencing by synthesis known as CoolMPS™. It uses antibodies that bind natural untagged nucleotides as they are incorporated. These antibodies are fluorescently labeled and are much cheaper to produce than labeled nucleotides, reducing the cost and increasing sequencing accuracy (Drmanac et al., 2020).
Scalability of new Illumina and MGI genetic sequencers
MGI sequencing platforms are the most scalable option for large-scale studies. For example, the DNBSEQ-T7 MGI genetic sequencer can produce up to 6 Tb of data, and 20 billion sequencing reads from four flow cells (MGI, 2023).
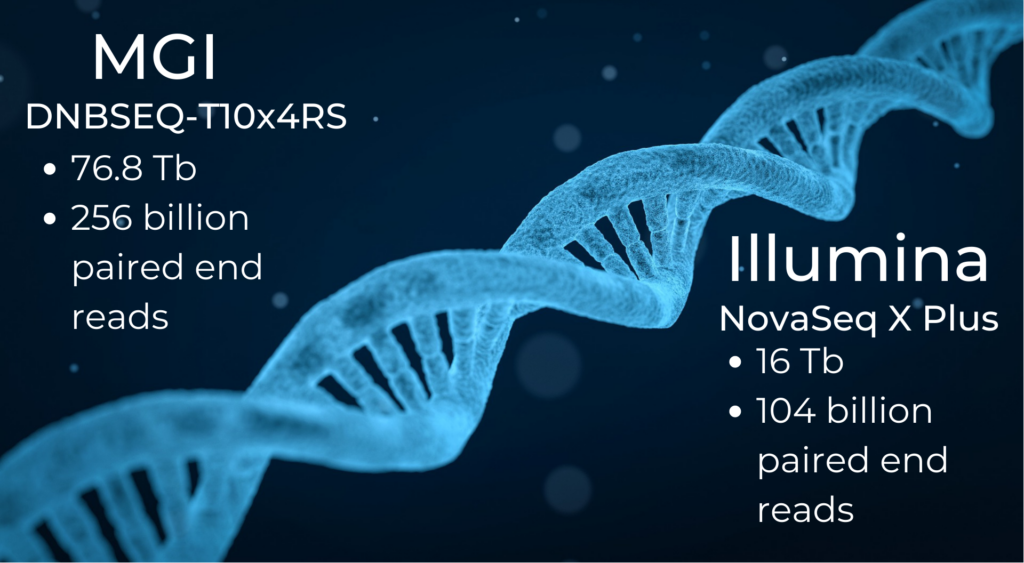
For ultra-high-depth whole genome sequencing, the MGI DNBSEQ-T10x4RS can produce a staggering 76.8 Tb of data per run and around 256 billion paired-end reads of 150 bp length from eight flow cells (MGI, 2023).
The highest throughput Illumina sequencing machine is the new NovaSeq X Plus. It has a maximum output of 16 Tb per run and can generate 104 billion paired-end reads (Illumina, 2023).
The increases in throughput of Illumina and MGI sequencing machines also allow more extensive multiplexing at deeper sequencing depths per sample. This means significant time and cost savings for researchers.
In the rapidly evolving world of genomic sequencers, future advances in throughput will undoubtedly enable higher scalability than ever before.
3’ mRNA-seq methods further increase scalability
A novel 3’ mRNA-seq library preparation method called MERCURIUS™ BRB-seq from Alithea Genomics is based on sample barcoding and tagging of 3’ ends of mRNA molecules. It is compatible with Illumina and MGI sequencing platforms.
With MERCURIUS™ BRB-seq, researchers can multiplex thousands of samples and produce similar quality data as conventional RNA-seq approaches, but with lower costs and hands-on time (Alpern et al., 2019).
When 3’ mRNA-seq technology is combined with ultra-high-throughput sequencing platforms, the number of samples researchers can process simultaneously is unparalleled.
Please contact us here to learn more about Illumina sequencing, MGI sequencing, or MERCURIUS™ BRB-seq.
References
- Alpern, D. et al. (2019) ‘BRB-seq: Ultra-affordable high-throughput transcriptomics enabled by bulk RNA barcoding and sequencing’, Genome Biology, 20(1), pp.1-15. Available at: https://doi.org/10.1186/s13059-019-1671-x.
- Drmanac, R. et al. (2010) ‘Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays’, Science, 327(5961), pp. 78–81. Available at: https://doi.org/10.1126/science.1181498.
- Drmanac, S. et al. (2020) ‘CoolMPS: Advanced massively parallel sequencing using antibodies specific to each natural nucleobase’. BioRxiv, pp.2020-02. Available at: https://doi.org/10.1101/2020.02.19.953307.
- Goodwin, S., McPherson, J.D. and McCombie, W.R. (2016) ‘Coming of age: Ten years of next-generation sequencing technologies’, Nature Reviews Genetics, 17(6), pp. 333–351pp. 333–351. Available at: https://doi.org/10.1038/nrg.2016.49.
- Illumina (2023) ‘Novaseq X plus’, Available at: https://www.illumina.com/systems/sequencing-platforms/novaseq-x-plus.html
- MGI (2023) ‘Sequencers’. Available at: https://en.mgi-tech.com/products/
- Mortazavi, A. et al. (2008) ‘Mapping and quantifying mammalian transcriptomes by RNA-Seq’, Nature Methods, 5(7), pp. 621–628. Available at: https://doi.org/10.1038/nmeth.1226.